True Covariance Matrix - \(\bb{\Sigma}\)
[,1] [,2] [,3]
[1,] 1.00 -0.26 0.31
[2,] -0.26 1.00 -0.08
[3,] 0.31 -0.08 1.00Network Psychometrics
Educational Statistics and Research Methods
Understand what is network psychometrics and
Understand the relationship between psychometric/psychological network with factor analysis
Understand how network model can be applied into real scenarios
Network analysis is a broad area. It has many names in varied fields:
1 and 2 focus on the probabilistic relationships and further casual relationships among variables. 3 and 4 focuses on network structure and node importance. 5 focus on the regression coefficients of structural and measurement mdoel.
All 5 analysis methods have a network-shaped diagram. Graphical modeling is a more “general” term that can comprise of the other network models.
Factor analysis (common factor model) assumes the associations between observed features can be explained by one or more common factors.
Psychometric network, however, assumes the associations between observed features ARE the reason of the development of depression. Or “depression” is the network itself.
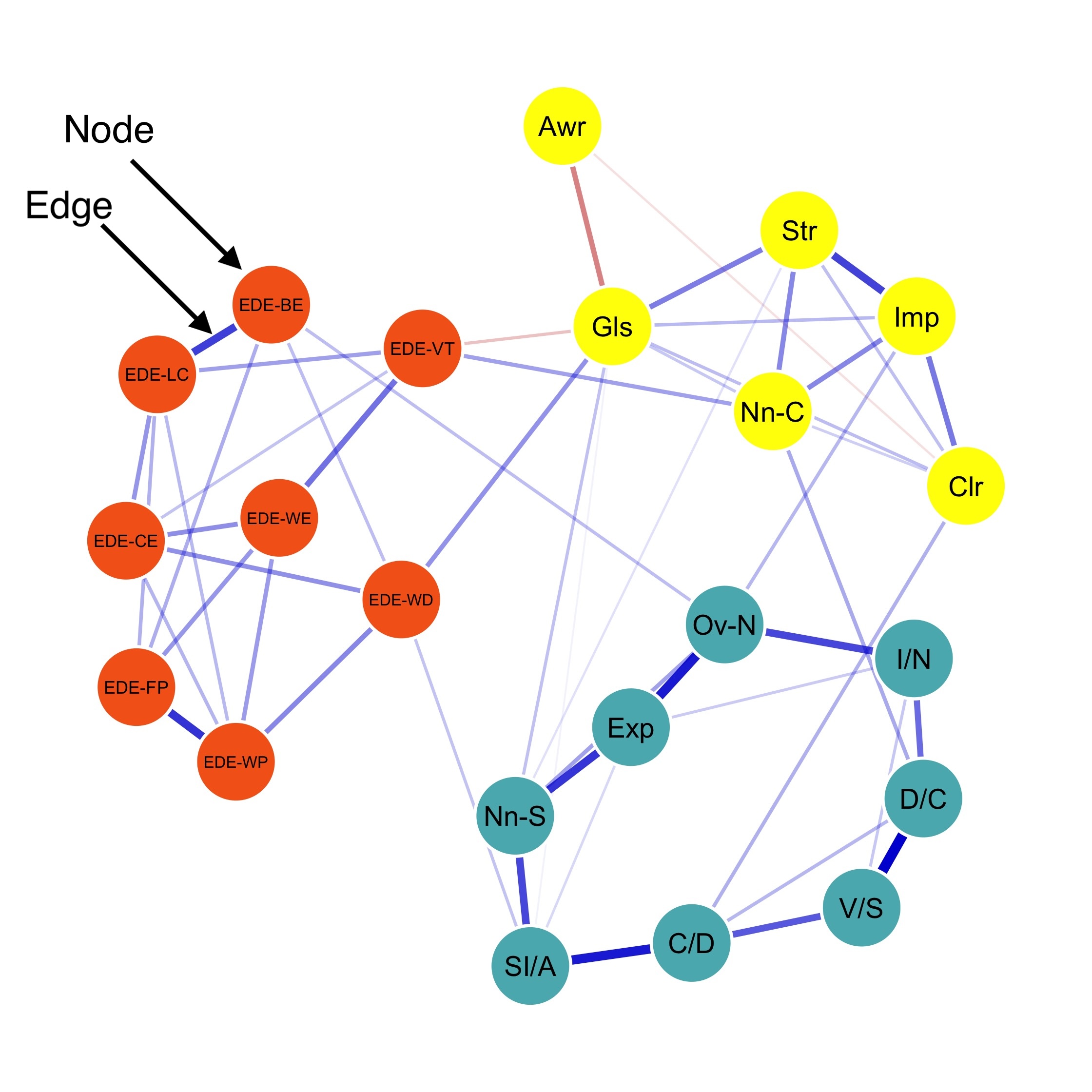
Pyschometric network is exploratory by nature. To obtain a meaningful network structure, psychometric networks need to drop weak edges but keep strong edges.
This procedure is typically called edge selection. One popular edge selection method is regularization.
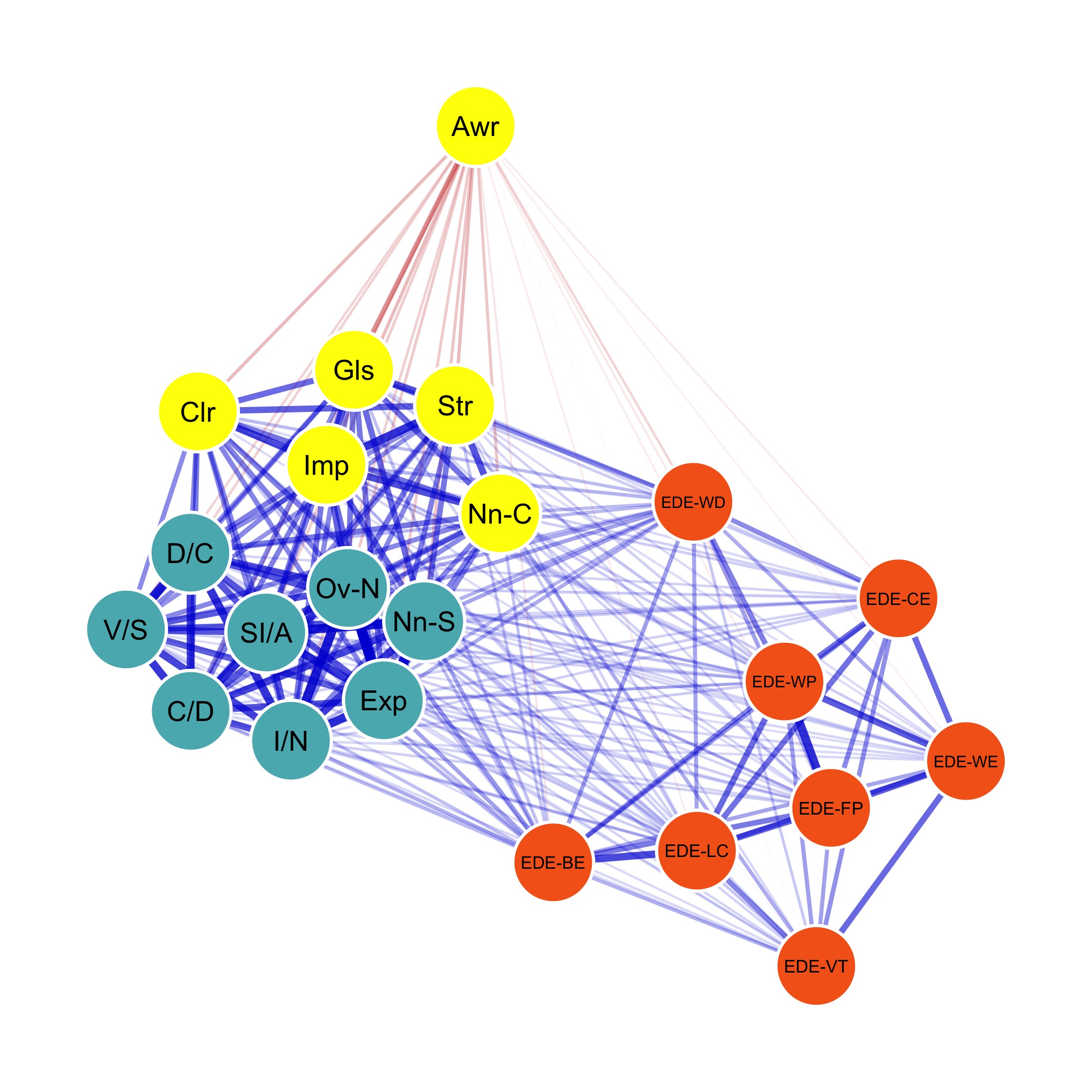


Psychometric network analysis methodology includes steps of network structure estimation (to construct the network), network description (to characterize the network) and network stability analysis (to assess the robustness of results).
GGM is one type of Pairwise Markov random field (PMRF) when data are continuous.
In a PMRF, the joint likelihood of multivariate data is modelled through the use of pairwise conditional associations, leading to a network representation that is undirected.
For \(p\)-dimensional data following multivariate normal distribution:
\[ \newcommand{\bb}[1]{\boldsymbol{#1}} \bb{X} \sim \mathcal{N}(\mu, \bb{K}^{-1}) \]
Where \(K\) is a inverse covariance matrix of \(\bb{X}\) (\(K = \Sigma^{-1}\)), also known as precision/concentration matrix.
To obtain sparse network structure, the \(i\)th row and \(j\)th column element of \(\bb{K}\), \(k_{ij}=0\) when edge \(\{j, k\}\) is not included in the network \(G\),
GGM can be standardized as the partial correlation network, in which each edge of GGM representing partial correlations between two nodes.
\[ \rho_{ij}=\rho(X^{(i)}, X^{(j)}|\bb{X}^{-(i,j)}) = -\frac{k_{ij}}{\sqrt{k_{ii}}\sqrt{k_{jj}}} \]
Assume there are three variables: fatigue, insomnia, concentration
[,1] [,2] [,3]
[1,] 1.00 -0.26 0.31
[2,] -0.26 1.00 -0.08
[3,] 0.31 -0.08 1.00Factor analysis model:
\[ \bb{X}\sim\mathcal{N}(\mu, \bb{\Lambda\Psi\Lambda^\text{T}+\Phi}) \]
GGM with partial correlation matrix:
\[ \bb{X} \sim \mathcal{N}(0, \bb{\Delta(I-\Omega)^{-1}\Delta}) \]
Where
psychonetrics: Partial correlation matrix estimation [,1] [,2] [,3]
[1,] 1.00 -0.26 0.31
[2,] -0.26 1.00 -0.08
[3,] 0.31 -0.08 1.00This is psychonetrics 0.13.2! For questions, issues, and bug reports, please see github.com/SachaEpskamp/psychonetrics.
Attaching package: 'psychonetrics'The following object is masked from 'package:graphics':
identifyAssuming denominator n was used in covariance computation (covtype = 'ML'). [,1] [,2] [,3]
[1,] 0.000 -0.248 0.300
[2,] -0.248 0.000 0.001
[3,] 0.300 0.001 0.000 [,1] [,2] [,3]
[1,] 0.921 0.000 0.000
[2,] 0.000 0.966 0.000
[3,] 0.000 0.000 0.951BGGM: Bayesian approachRegistered S3 methods overwritten by 'ergm':
method from
simulate.formula lme4
simulate.formula_lhs lme4
summary.formula HmiscRegistered S3 methods overwritten by 'BFpack':
method from
get_estimates.lm bain
get_estimates.t_test bainRegistered S3 method overwritten by 'GGally':
method from
+.gg ggplot2
Attaching package: 'BGGM'The following object is masked from 'package:psychonetrics':
precisionBGGM: Posterior Sampling BGGM: Finished [,1] [,2] [,3]
[1,] 0.000 -0.260 0.189
[2,] -0.260 0.000 0.042
[3,] 0.189 0.042 0.000BGGM: Bayesian Gaussian Graphical Models
---
Type: continuous
Analytic: FALSE
Formula:
Posterior Samples: 1000
Observations (n):
Nodes (p): 3
Relations: 3
---
Call:
BGGM::estimate(Y = dat, type = "continuous", analytic = FALSE,
iter = 1000)
---
Estimates:
Relation Post.mean Post.sd Cred.lb Cred.ub
1--2 -0.260 0.042 -0.342 -0.178
1--3 0.189 0.043 0.104 0.272
2--3 0.042 0.047 -0.052 0.129
--- Multiple procedure and software can be used to perform edge selection:
prune function in psychonetrics package uses stepdown model search by pruning non-significant parameters
EBICglasso in qgraph and glasso package uses Extended Bayesian Information Criterion (EBIC) to select best model
\[ \text{EBIC}=-2\text{L}+E(\log(N))+4\gamma E(log(P)) \]
select function in BGGM package uses Bayesian Hypothesis Testing — Bayes Factor (BF) to select model
25 self-reported personality items representing 5 factors.
70.6% edges are nonzero using EBICglasso, while only 63.6% edges are nonzero using BF method and 62.0% edges are kept using \(\alpha =.01\) significance testing
Attaching package: 'psych'The following object is masked from 'package:BGGM':
bfiThe following object is masked from 'package:psychonetrics':
bifactorVariables detected as ordinal: A1; A2; A3; A4; A5; C1; C2; C3; C4; C5; E1; E2; E3; E4; E5; N1; N2; N3; N4; N5; O1; O2; O3; O4; O5EBICgraph <- EBICglasso(CorMat, nrow(bfi), 0.5, threshold = TRUE)
## density
density_nonzero_edge <- function(pcor_matrix){
N_nonzero_edge = (sum(pcor_matrix == 0) - ncol(pcor_matrix)) /2
N_all_edge = ncol(pcor_matrix)*(ncol(pcor_matrix)-1)/2
N_nonzero_edge/N_all_edge
}
density_nonzero_edge(EBICgraph)[1] 0.7066667[1] 0.62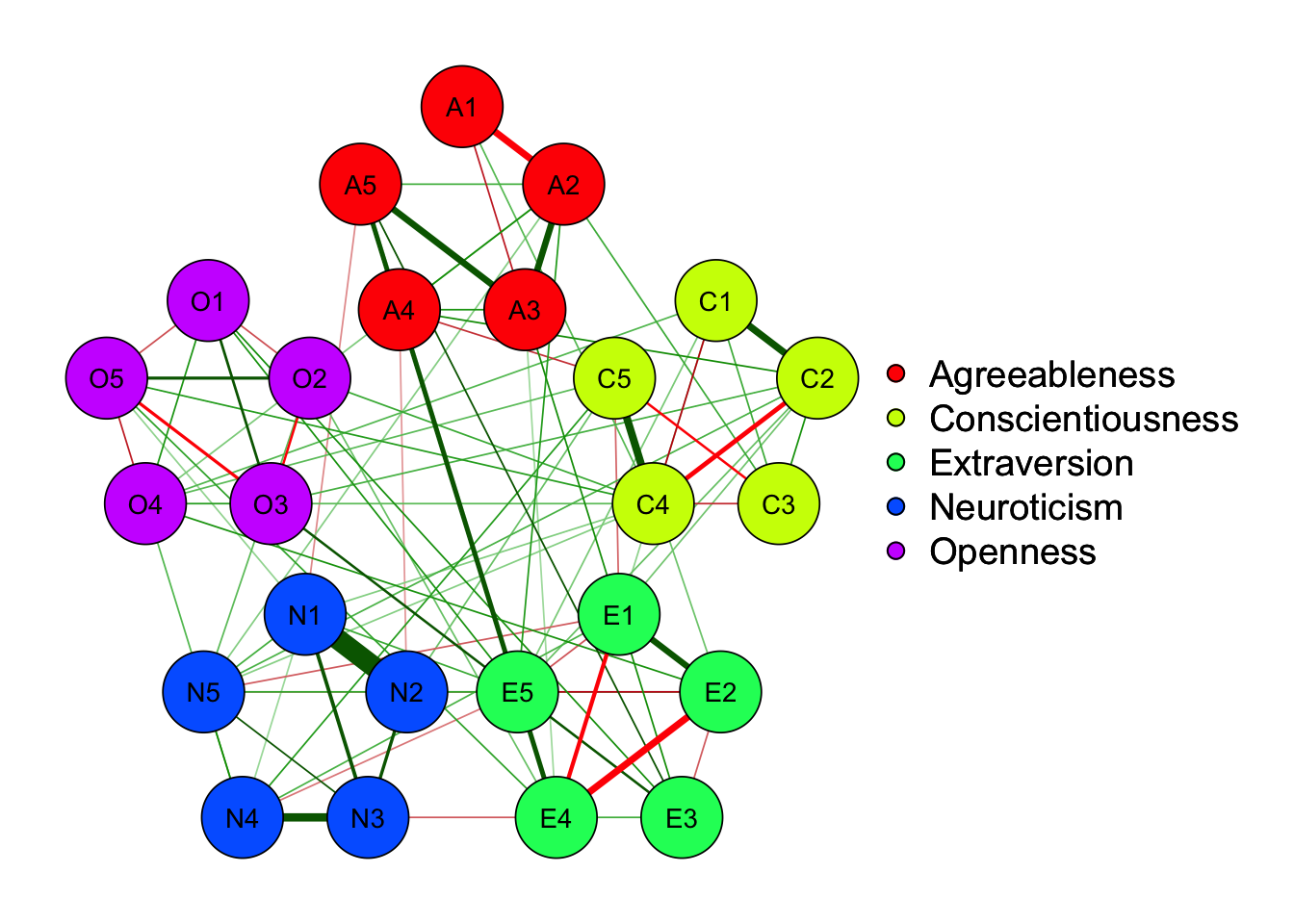
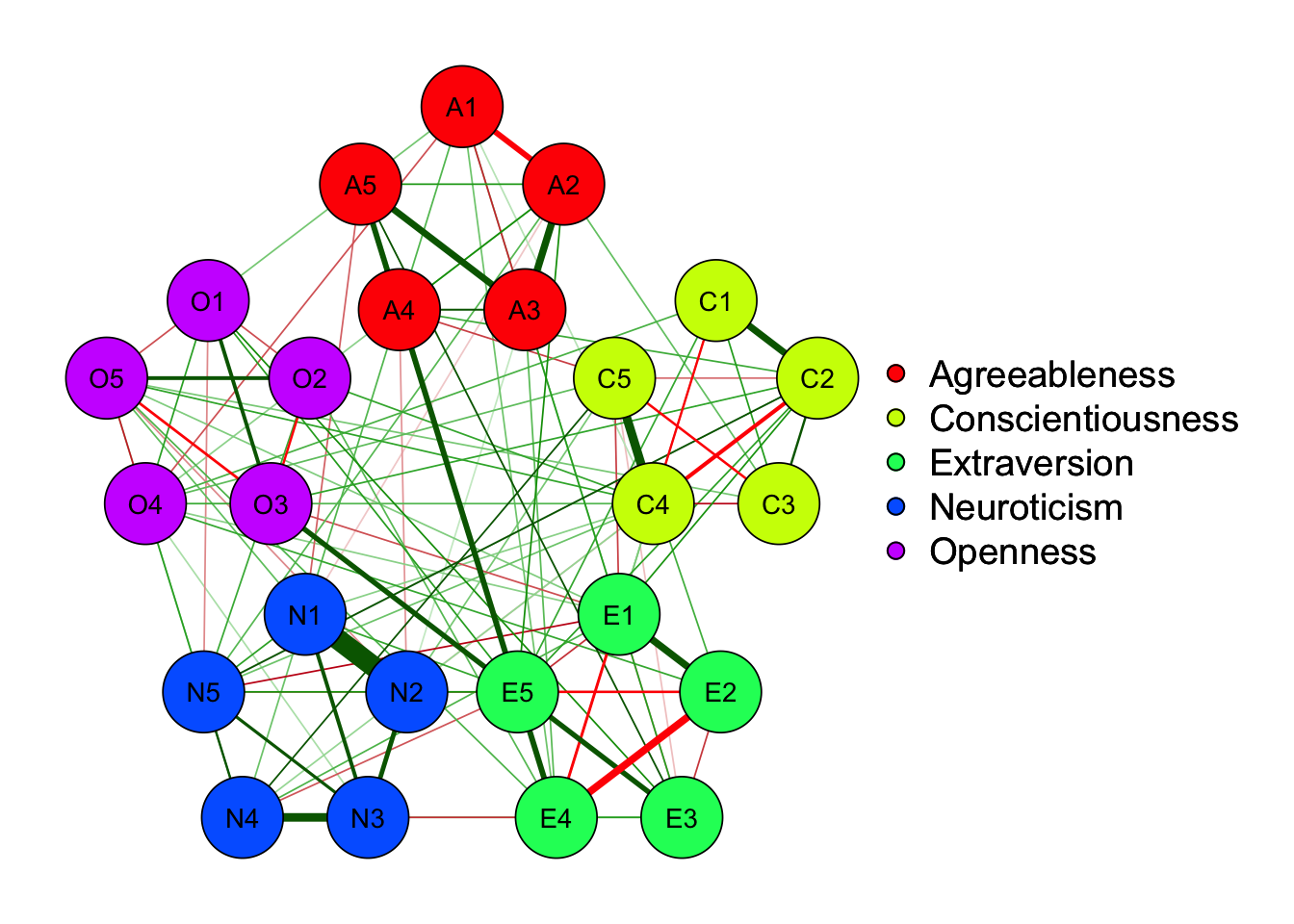
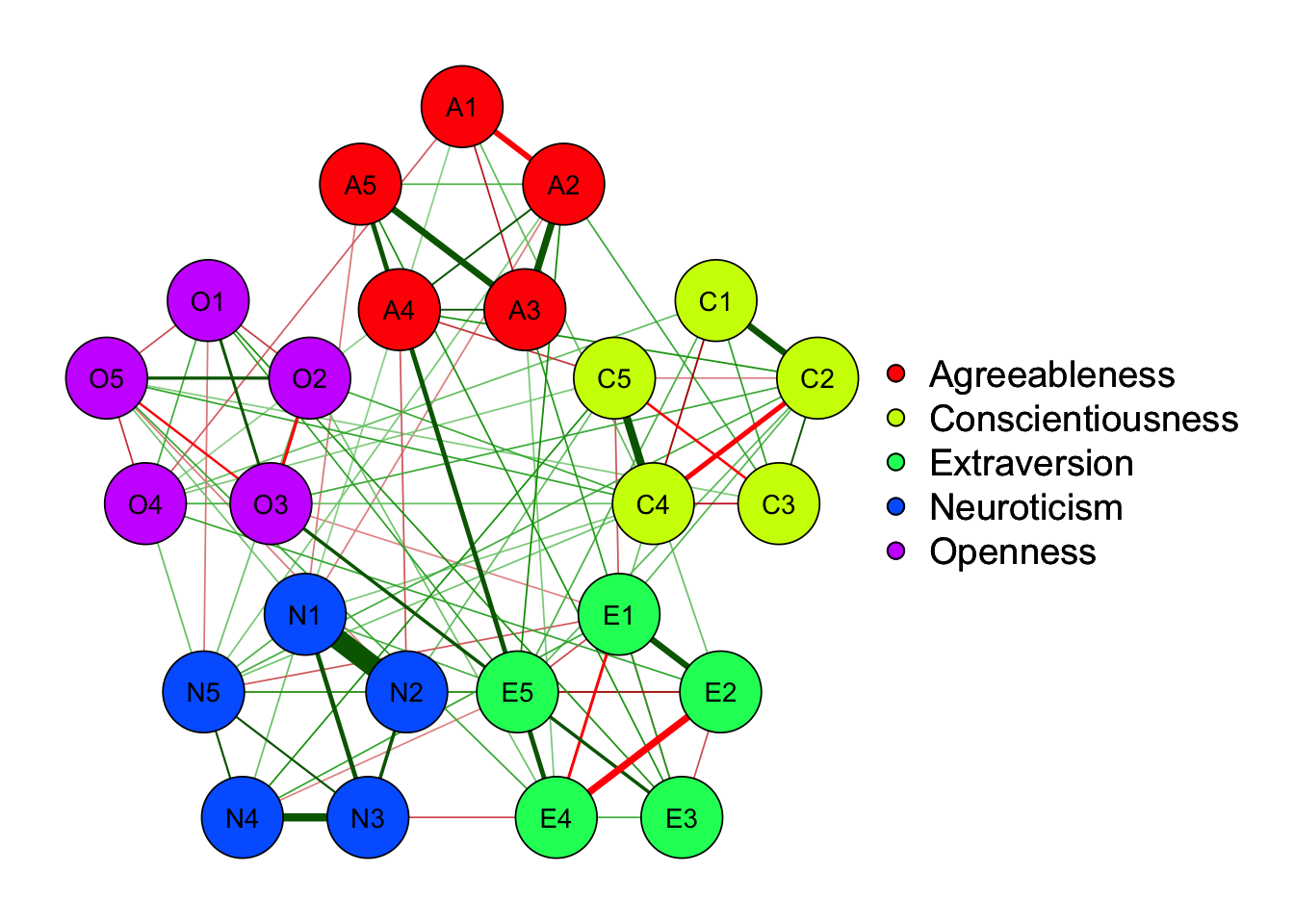
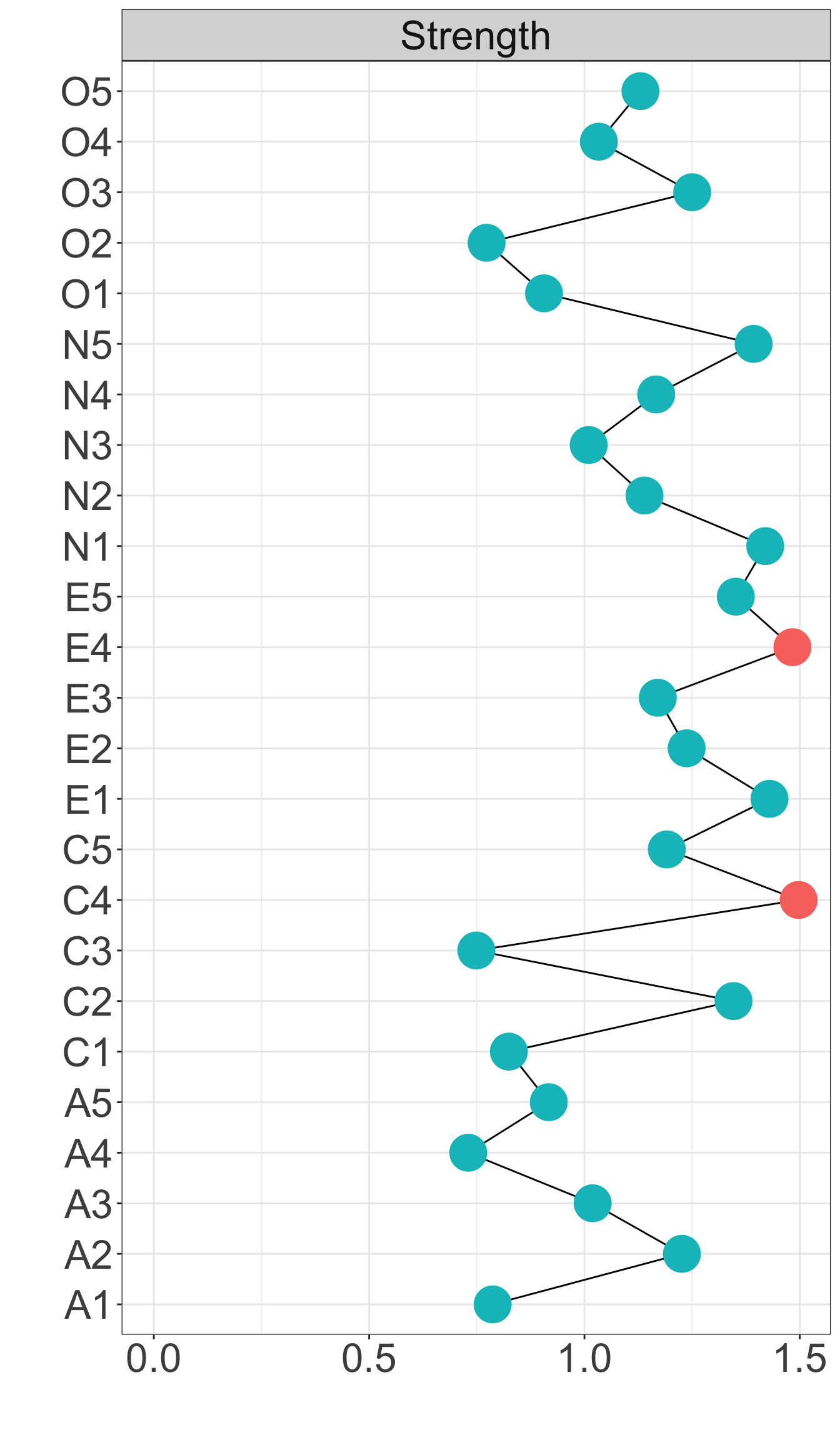
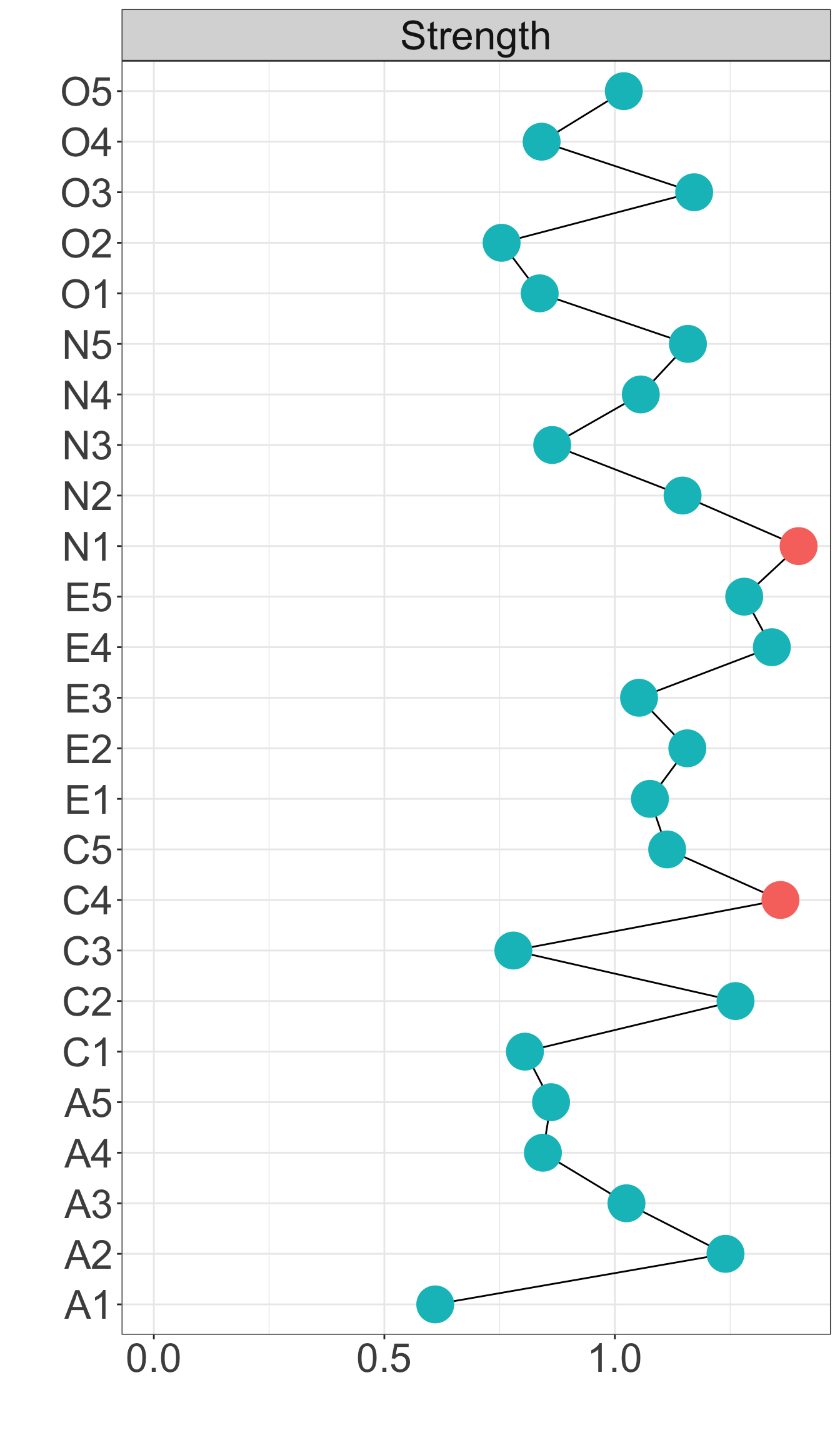
Strength centrality measures suggest that C4 and E4 or N1 have highest centrality indicating they play most imporatant roles in the networks.
Attaching package: 'ggplot2'The following objects are masked from 'package:psych':
%+%, alpha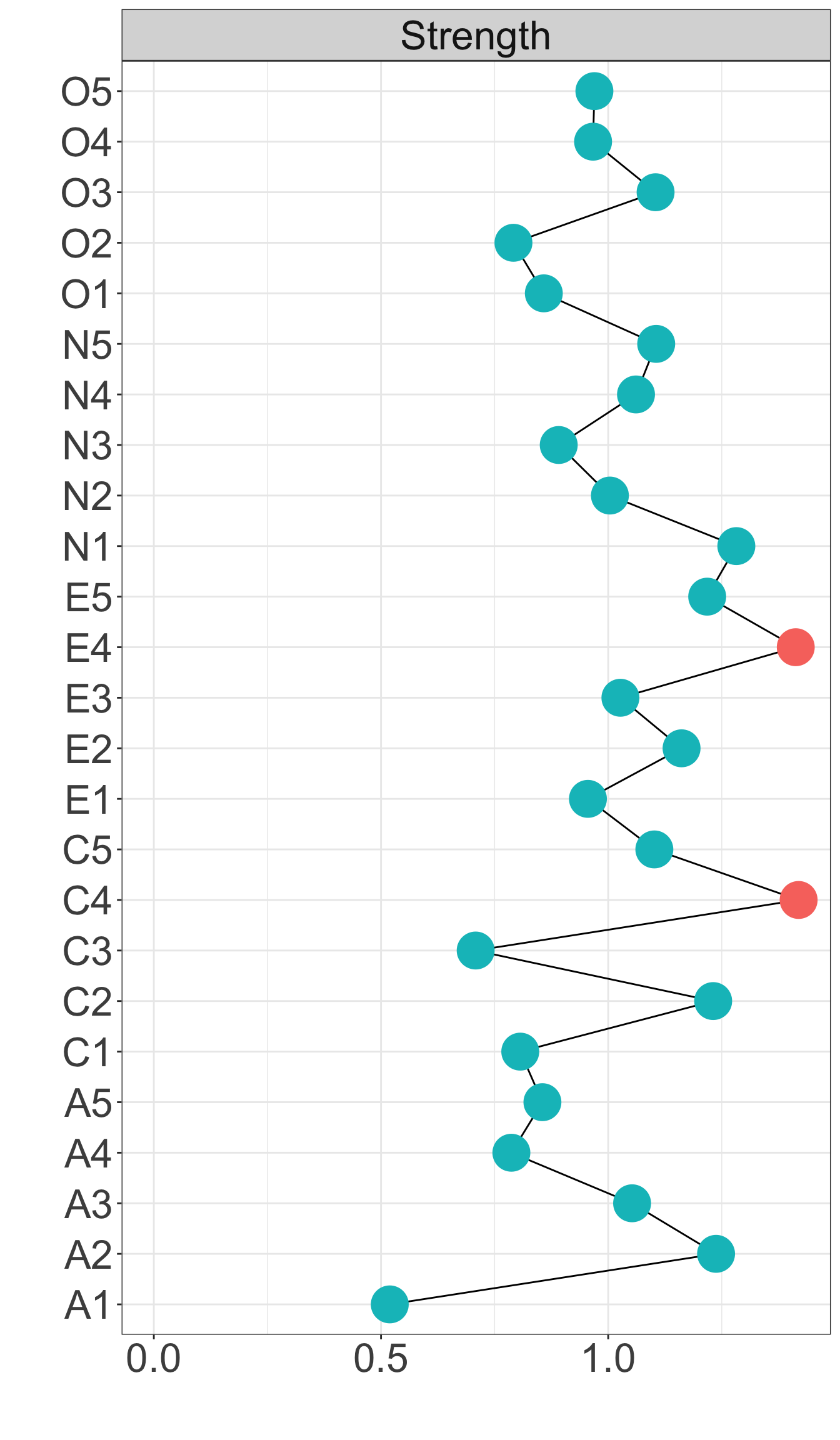
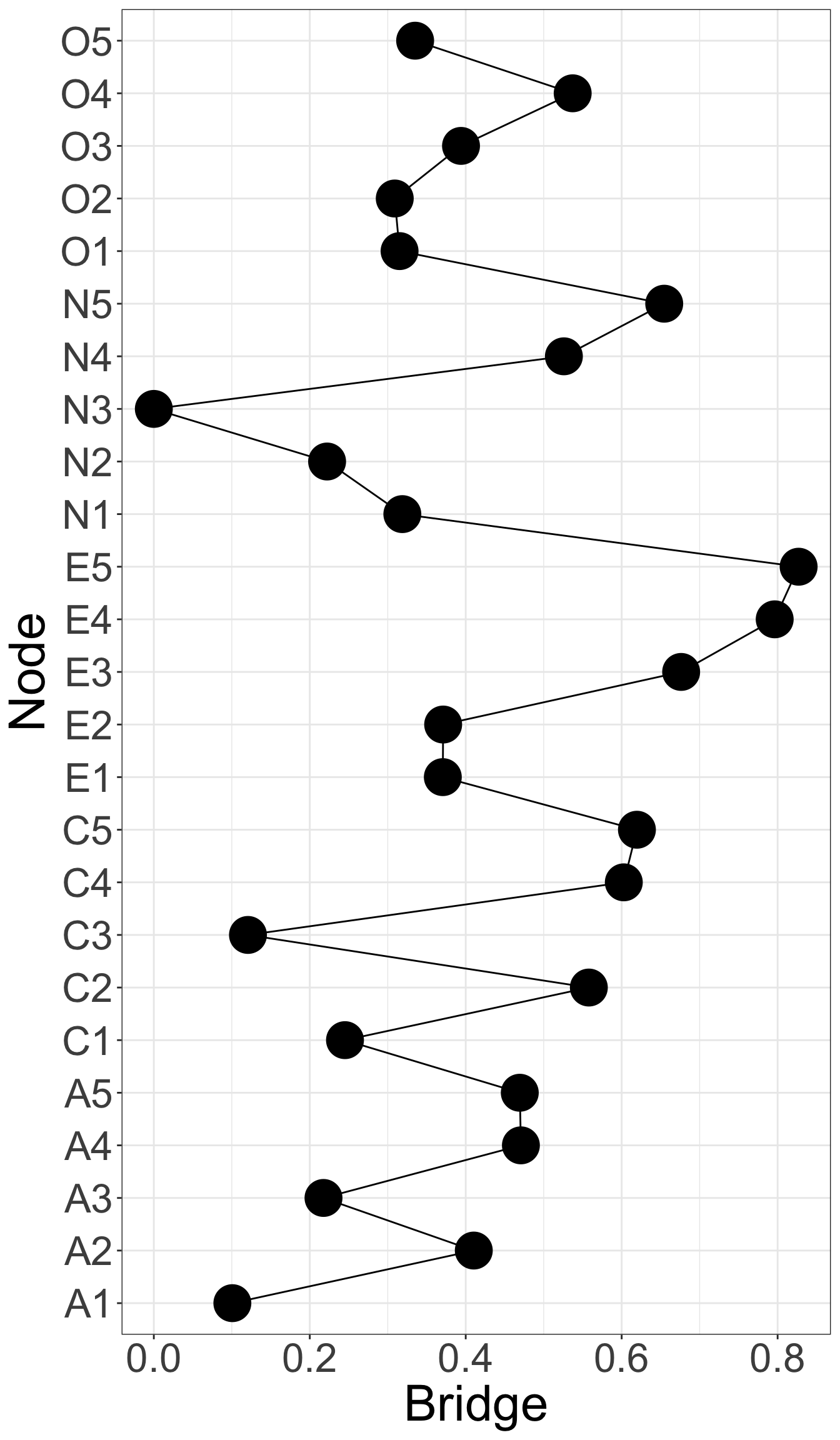
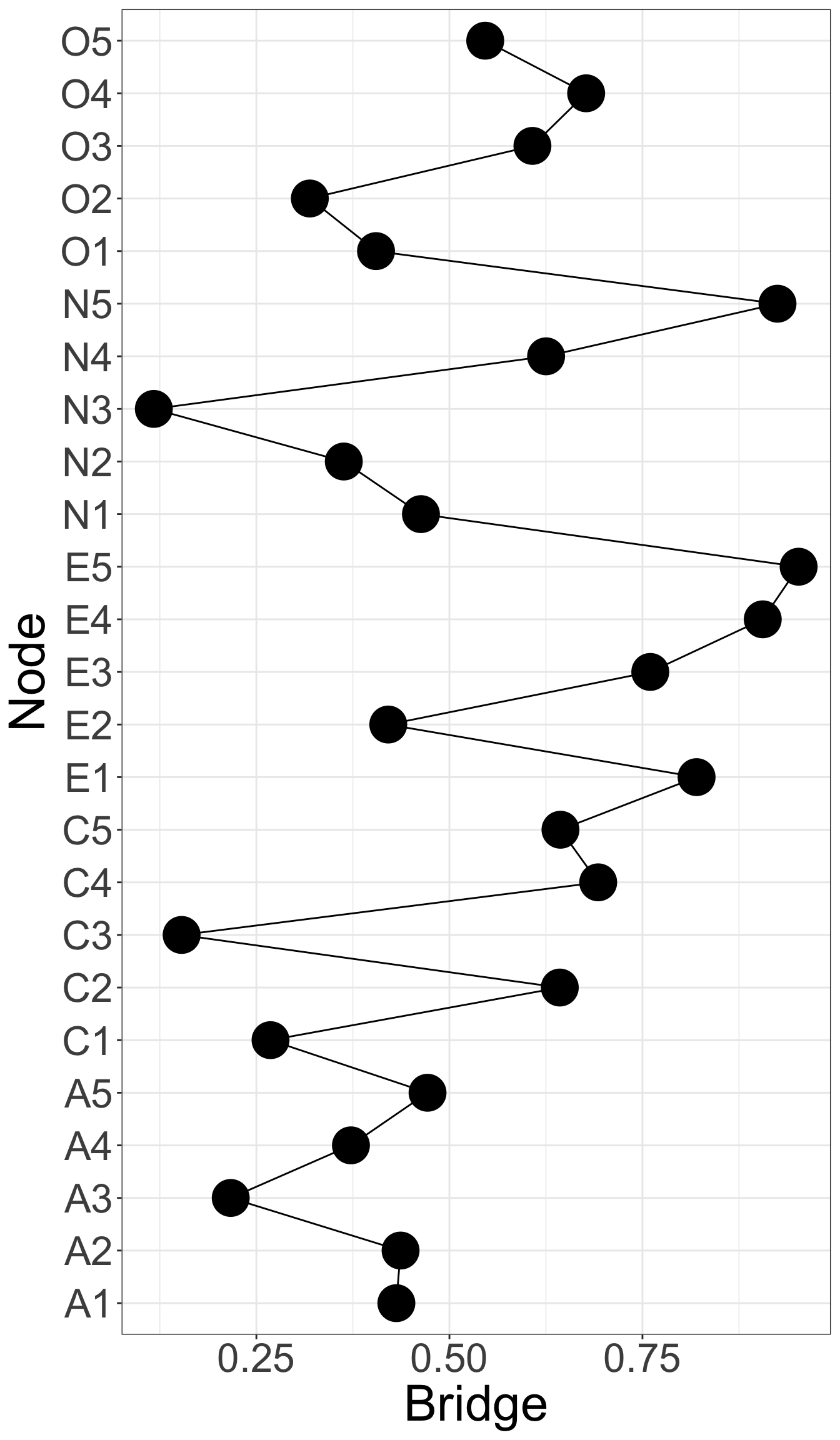
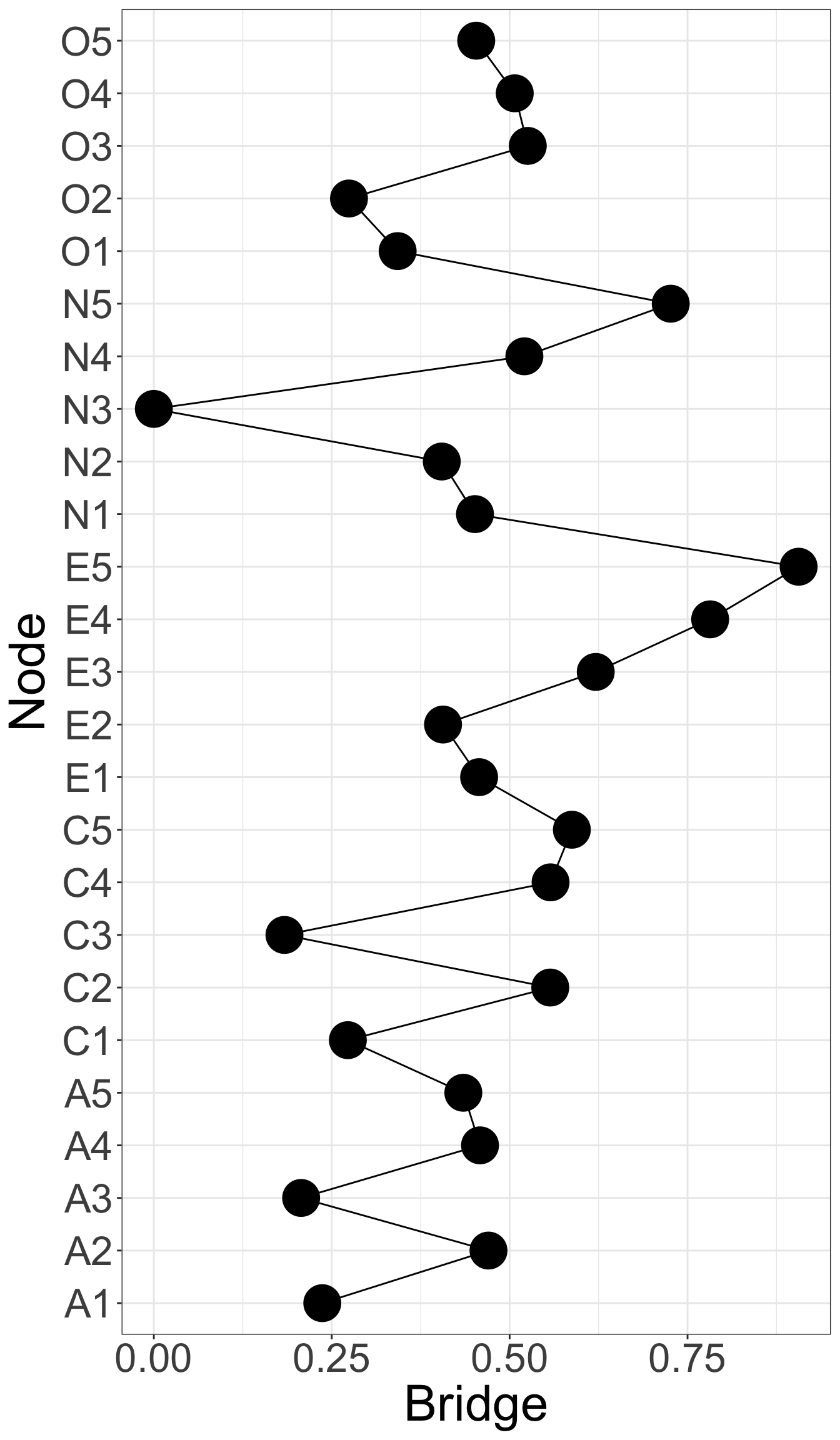
E4 and E5 has highest bridge strength, indicating they serve as bridges linking communities of personality. They are important elements connecting varied types of personality
── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ dplyr 1.1.4 ✔ readr 2.1.5
✔ forcats 1.0.0 ✔ stringr 1.5.1
✔ lubridate 1.9.4 ✔ tibble 3.2.1
✔ purrr 1.0.4 ✔ tidyr 1.3.1
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ ggplot2::%+%() masks psych::%+%()
✖ ggplot2::alpha() masks psych::alpha()
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
✖ purrr::map() masks BGGM::map()
✖ dplyr::select() masks BGGM::select()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors
Registered S3 methods overwritten by 'networktools':
method from
print.bridge bridgesampling
summary.bridge bridgesampling