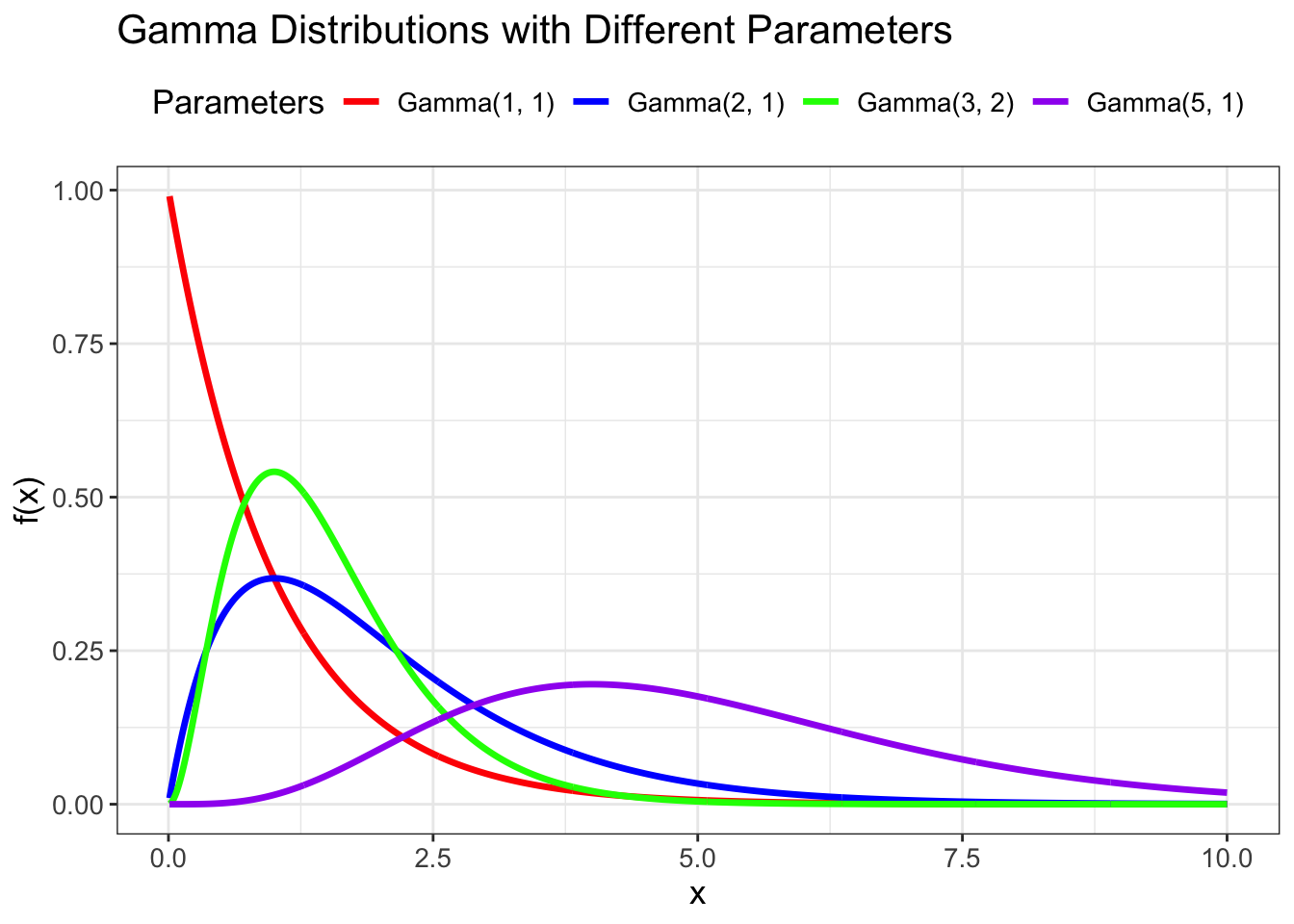
# Gamma distribution with different shape and rate parameters
x <- seq(0.01, 10, .01)
shape_params <- c(1, 2, 3, 5)
rate_params <- c(1, 1, 2, 1)
# Create data for different gamma distributions
gamma_data <- data.frame()
for(i in 1:length(shape_params)) {
temp_data <- data.frame(
x = x,
density = dgamma(x, shape = shape_params[i], rate = rate_params[i]),
distribution = paste0("Gamma(", shape_params[i], ", ", rate_params[i], ")")
)
gamma_data <- rbind(gamma_data, temp_data)
}
# Plot the distributions
ggplot(gamma_data) +
geom_path(aes(x = x, y = density, color = distribution), linewidth = 1.2) +
scale_color_manual(values = c("red", "blue", "green", "purple")) +
labs(x = "x", y = "f(x)",
title = "Gamma Distributions with Different Parameters",
color = "Parameters") +
theme_bw() +
theme(legend.position = "top", text = element_text(size = 13))