---
title: "Lecture 07"
subtitle: "Generalized Measurement Models: Modeling Observed Data"
author: "Jihong Zhang"
institute: "Educational Statistics and Research Methods"
title-slide-attributes:
data-background-image: ../Images/title_image.png
data-background-size: contain
execute:
echo: true
eval: false
format:
html:
code-tools: true
code-line-numbers: false
code-fold: show
code-summary: 'Click here to fold R code'
highlight-style: arrow
number-offset: 1
fig.width: 10
fig-align: center
message: false
revealjs:
output-file: slides-index.html
logo: ../Images/UA_Logo_Horizontal.png
incremental: false # choose "false "if want to show all together
transition: slide
background-transition: fade
theme: [simple, ../pp.scss]
footer: <https://jihongzhang.org/posts/2024-01-12-syllabus-adv-multivariate-esrm-6553>
scrollable: true
slide-number: true
chalkboard: true
number-sections: false
code-line-numbers: true
code-annotations: below
code-copy: true
code-summary: ''
highlight-style: arrow
view: 'scroll' # Activate the scroll view
scrollProgress: true # Force the scrollbar to remain visible
mermaid:
theme: neutral
#bibliography: references.bib
---
## Previous Class
1. We used two simulation data sets to introduce how Stan can be used for factor analysis
- A simple structure
- A structure with cross-loading
2. What we left is how to interpret parameters in factor analysis
3. We also need to refresh our memory of R coding and Stan coding we've learnt so far
## Today's Lecture Objectives
1. Quickly go through our R code file so far
2. Show different modeling specifications for different types of item response data
3. Show how parameterization differs for standardized latent variables vs. marker item scale identification
::: {.callout-note appearance="minimal"}
**Lecture 07 Files:** [{{< fa file-code >}} Lecture07.R](Code/Lecture07.R){download="Lecture07.R"} | [{{< fa file-csv >}} conspiracies.csv](Code/conspiracies.csv){download="conspiracies.csv"}
:::
## Example Data: Conspiracy Theories
- Today's example is from a bootstrap resample of 177 undergraduate students at a large state university in the Midwest.
- The survey was a measure of 10 questions about their beliefs in various conspiracy theories that were being passed around the internet in the early 2010s
- All item responses were on a 5-point Likert scale with:
- Strong Disagree
- Disagree
- Neither Agree nor Disagree
- Agree
- Strongly Agree
- The purpose of this survey was to study individual beliefs regarding conspiracies.
- Our purpose in using this instrument is to provide a context that we all may find relevant as many of these conspiracies are still prevalent.
## Conspiracy Theory Q1-Q5[^1]
[^1]: Built by Dr. Jonathan Templin
1. The U.S. invasion of Iraq was not part of a campaign to fight terrorism, but was driven by oil companies and Jews in the U.S. and Israel.
2. Certain U.S. government officials planned the attacks of September 11, 2001 because they wanted the United States to go to war in the Middle East.
3. President Barack Obama was not really born in the United States and does not have an authentic Hawaiian birth certificate.
4. The current financial crisis was secretly orchestrated by a small group of Wall Street bankers to extend the power of the Federal Reserve and further their control of the world's economy.
5. Vapor trails left by aircraft are actually chemical agents deliberately sprayed in a clandestine program directed by government officials.
## Conspiracy Theory Q6-Q10
1. Billionaire George Soros is behind a hidden plot to destabilize the American government, take control of the media, and put the world under his control.
2. The U.S. government is mandating the switch to compact fluorescent light bulbs because such lights make people more obedient and easier to control.
3. Government officials are covertly Building a 12-lane "NAFTA superhighway" that runs from Mexico to Canada through America's heartland.
4. Government officials purposely developed and spread drugs like crack-cocaine and diseases like AIDS in order to destroy the African American community.
5. God sent Hurricane Katrina to punish America for its sins.
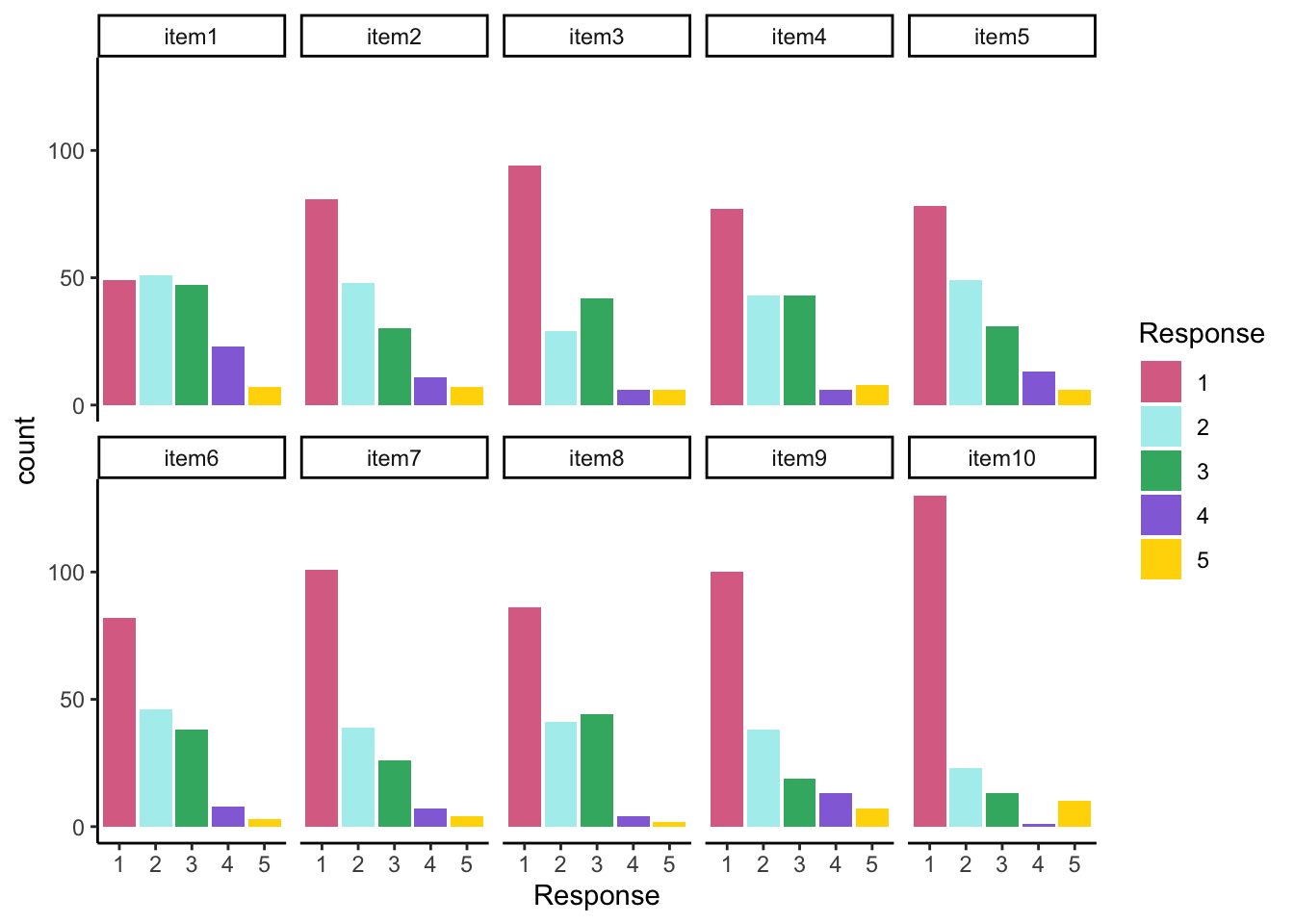
## Response Distribution
All items seem to be positive skewed
```{r}
#| eval: true
#| code-fold: true
#| collapse: true
#| warning: false
#| message: false
library(tidyverse)
library(here)
self_color <- c("#DB7093", "#AFEEEE", "#3CB371", "#9370DB", "#FFD700")
root_dir <- "teaching/2024-01-12-syllabus-adv-multivariate-esrm-6553/Lecture07/Code"
dat <- read.csv(here(root_dir, 'conspiracies.csv'))
itemResp <- dat[,1:10]
colnames(itemResp) <- paste0('item', 1:10)
conspiracyItems = itemResp
itemResp |>
rownames_to_column("ID") |>
pivot_longer(-ID, names_to = "Item", values_to = "Response") |>
mutate(Item = factor(Item, levels = paste0('item', 1:10)),
Response = factor(Response, levels = 1:5)) |>
ggplot() +
geom_bar(aes(x = Response, fill = Response, group = Response),
position = position_stack()) +
facet_wrap(~ Item, nrow = 2, ncol = 5) +
theme_classic() +
scale_fill_manual(values = self_color)
```
## Conspiracy Theories: Assumed Latent Variable
For today's lecture, we will assume each of the 10 items measures one single latent variable
$\theta$: tendency to believe in conspiracy theories
- Higher value of $\theta$ suggests more likelihood of believing in conspiracy theories
- Let's denote this latent variable as $\theta_p$ for individual *p*
- *p* is the index for person with $p = \{1, \cdots, P\}$
- We will assume this latent variable is:
- Continuous
- Normally distribution: $\theta_p \sim N(\mu_\theta, \sigma_\theta)$
- We will make different assumptions about the response distribution to show how prior settings affect results
- Across all people, we will denote the set of vector of latent variable as
$$
\Theta = \begin{bmatrix}\theta_1, \cdots, \theta_P\end{bmatrix}^T
$$
---
{width="100%" fig-alt="CFA path diagram showing latent variable theta (Conspiracy Belief) with factor loadings to all 10 observed items Q1–Q10, each with an intercept and unique variance."}
# Building Measurement Model
## Observed Variables with Normal Distributions
A psychometric model posits that one or more hypothesized latent variable(s) is the common cause that can predict a person's response to observed items:
1. Our hypothesized latent variable: Tendency to Believe in Conspiracies ($\theta_p$)
2. As we have only one variable, the model structure is called `Unidimensional`
3. All 10 items are considered as outcomes of the latent variable in the model
4. In today's class, we assume all item response follow a normal distribution:
- This is the assumption underlying confirmatory factor analysis (CFA) models
- This assumption is tenuous at best
## Normal Distribution: Linear Regression
A typical linear regression is like
$$
Y_p =\beta_0 +\beta_1 X_p + e_p
$$
with $e_p\sim N(0, \sigma_e)$
If we replace $X_p$ with latent variable $\theta_p$, and replace $\beta$ as factor loading $\lambda$
We can get the linear regression function (**IRF**) for each item
---
{width="100%" fig-alt="Diagram showing the term-by-term transformation from linear regression to the CFA item response function, with color-coded mapping of intercept, slope, predictor, and residual to their CFA equivalents."}
$$
Y_{p1} =\mu_{0} +\lambda_{1} \theta_p + e_{p1}; \ \ \ \ \ e_{p1}\sim N(0, \psi_1^2) \\
Y_{p2} =\mu_{2} +\lambda_{2} \theta_p + e_{p2}; \ \ \ \ \ e_{p2}\sim N(0, \psi_{2}^2) \\
Y_{p3} =\mu_{3} +\lambda_{3} \theta_p + e_{p3}; \ \ \ \ \ e_{p3}\sim N(0, \psi_{3}^2) \\
Y_{p4} =\mu_{4} +\lambda_{4} \theta_p + e_{p4}; \ \ \ \ \ e_{p4}\sim N(0, \psi_{4}^2) \\
Y_{p5} =\mu_{5} +\lambda_{5} \theta_p + e_{p5}; \ \ \ \ \ e_{p5}\sim N(0, \psi_{5}^2) \\
Y_{p6} =\mu_{6} +\lambda_{6} \theta_p + e_{p6}; \ \ \ \ \ e_{p6}\sim N(0, \psi_{6}^2) \\
Y_{p7} =\mu_{7} +\lambda_{7} \theta_p + e_{p7}; \ \ \ \ \ e_{p7}\sim N(0, \psi_{7}^2) \\
Y_{p8} =\mu_{8} +\lambda_{8} \theta_p + e_{p8}; \ \ \ \ \ e_{p8}\sim N(0, \psi_{8}^2) \\
Y_{p9} =\mu_{9} +\lambda_{9} \theta_p + e_{p9}; \ \ \ \ \ e_{p9}\sim N(0, \psi_{9}^2) \\
Y_{p10} =\mu_{10} +\lambda_{10} \theta_p + e_{p10}; \ \ \ \ \ e_{p10}\sim N(0, \psi_1{0}^2) \\
$$
## Interpretation of Parameters
- $\mu_i$: Item intercept
- Interpretation: the expected score on the item $i$ when $\theta_p=0$
- Higher Item intercept suggests more likely to believe in conspiracy for people with average level of conspiracy belief
- So it is also called **item easiness** in item response theory (IRT)
- $\lambda_i$: Factor loading or Item discrimination
- The change in the expected score of an item for a one-unit increase in belief in conspiracy
- $\psi_i^2$: Unique variance[^2]
[^2]: In **Stan**, we will specify $\psi_e$: the unique standard deviation
## Measurement Model Identification
When we specify measurement model, we need to choose on scale identification method for latent variable
1. Assume latent variable is normal distribution
2. Or, maker item has factor loading as "1"
In this study, we assume $\theta_p \sim N(0,1)$ which allows us to estimate all item parameters of the model
- This is what we call a **standardization identification method**
- Factor scores are like Z-scores
---
{width="100%" fig-alt="Side-by-side comparison of two scale identification methods: Method 1 fixes theta to N(0,1) so all item parameters are free; Method 2 fixes lambda-1 to 1 so theta's scale is determined by item 1."}
## Implementing Normal Outcomes in Stan
Recall that we can use matrix operation to make Stan estimate psychometric models with normal outcomes:
- The model (predictor) matrix cannot be used
- This is because the latent variable will be sampled so that the model matrix cannot be formed as a constant
- The data will be imported as a matrix
- More than one outcome means more than one column vector of data
- The parameters will be specified as vectors of each type
- Each item will have its own set of parameters
- Implications for the use of prior distributions
## Stan's `data` Block
```{stan, output.var="display"}
data {
int<lower=0> nObs; // number of observations #<1>
int<lower=0> nItems; // number of items #<2>
matrix[nObs, nItems] Y; // item responses in a matrix
vector[nItems] meanMu;
matrix[nItems, nItems] covMu; // prior covariance matrix for coefficients #<3>
vector[nItems] meanLambda; // prior mean vector for coefficients
matrix[nItems, nItems] covLambda; // prior covariance matrix for coefficients #<4>
vector[nItems] psiRate; // prior rate parameter for unique standard deviations #<5>
}
```
1. `nObs` is 177, declared as integer with lower bound as 0
2. `nItems` is 11, declared as integer with lower bound as 0
3. `meanMu` as `covMu` are prior mean and covariance matrix for $\mu_i$
4. `meanLambda` and `covLambda` are prior mean and covariance matrix for $\lambda_i$
5. `psiRate` is prior rate parameter for $\psi_i$
## Stan's `parameter` Block
```{stan, output.var='display'}
parameters {
vector[nObs] theta; // the latent variables (one for each person)
vector[nItems] mu; // the item intercepts (one for each item)
vector[nItems] lambda; // the factor loadings/item discriminations (one for each item)
vector<lower=0>[nItems] psi; // the unique standard deviations (one for each item)
}
```
Here, the parameterization of $\lambda$ (factor loadings / item discrimination) can lead to problems in estimation
- The issue: $\lambda_i \theta_p = (-\lambda_i) (-\theta_p)$
- Depending on the random starting values of each of these parameters (per chain), a given chain may converge to a different region
- To demonstrate, we will start with different random number seed (`seed`)
- Currently using **09102022**: works fine
- Change to **25102022**: big problem
```{r}
#| eval: false
#| code-summary: "R file for running the model with different seed"
modelCFA_samples = modelCFA_stan$sample(
data = modelCFA_data,
seed = 09102022,
chains = 4,
parallel_chains = 4,
iter_warmup = 1000,
iter_sampling = 2000
)
```
## Stan's model Block
```{stan, output.var='display'}
model {
lambda ~ multi_normal(meanLambda, covLambda); // Prior for item discrimination/factor loadings
mu ~ multi_normal(meanMu, covMu); // Prior for item intercepts
psi ~ exponential(psiRate); // Prior for unique standard deviations
theta ~ normal(0, 1); // Prior for latent variable (with mean/sd specified)
for (item in 1:nItems){
Y[,item] ~ normal(mu[item] + lambda[item]*theta, psi[item]);
}
}
```
The loop here conducts the model via item response function (**IRF**) for each item:
- Assumption of conditional independence enables this
- Non-independence would need multivariate normal model
- The item mean is set by the conditional mean of the model
- The item SD is set by the unique variance parameter
- The loop puts each item's parameters into the question
## Choosing Prior Distributions for Parameters
There is not uniform agreement about the choices of prior distributions for item parameters
- We will use **uninformative** priors on each to begin
- After first model analysis, we will discuss these choices and why they were made
- For now:
- Item intercepts: $\mu_i \sim N(0, \sigma_{\mu_i}^2 = 1000)$
- Factor loadings / item discrimination: $\lambda_i \sim N(0, \sigma^2_{\lambda_i}=1000)$
- Unique standard deviations: $\psi_i \sim \text{exponential}(r_{\psi_i} = .01)$
## Prior Density Function Plots
::::: columns
::: {.column width="50%"}
Prior distribution for item intercepts, factor loadings
N(0, 1000)
```{r}
#| eval: true
#| echo: false
#| code-fold: true
#| fig-height: 8
set.seed(1234)
data.frame(
x = seq(0, 5, .001),
y = dnorm(x = seq(0, 5, .001), mean = 0, sd = sqrt(1000))
) |>
ggplot(aes(x=x, y =y)) +
geom_path(linewidth = 1.3) +
labs(x = "", y = "Probability") +
theme_classic() +
theme(text = element_text(size = 25))
```
:::
::: {.column width="50%"}
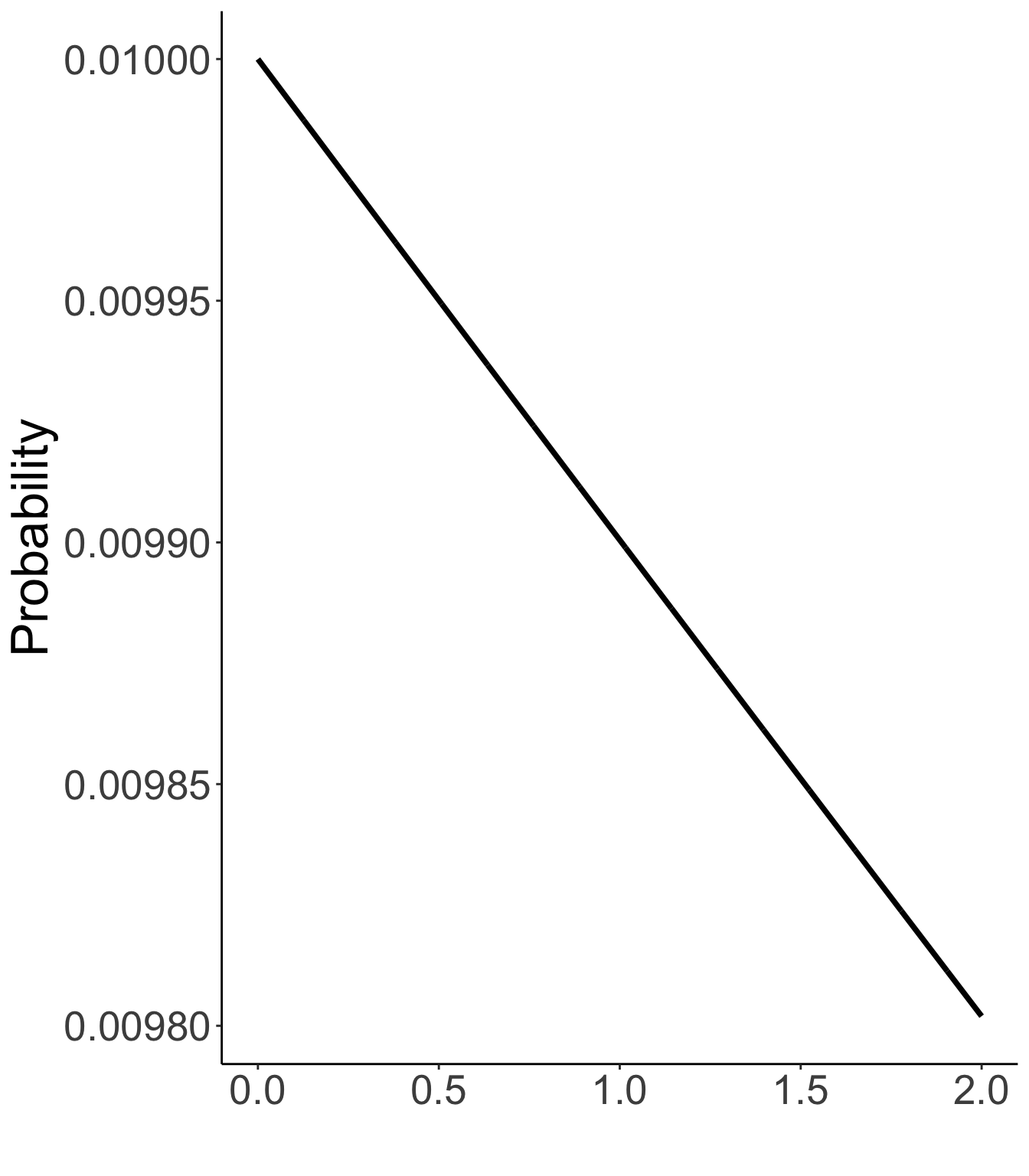
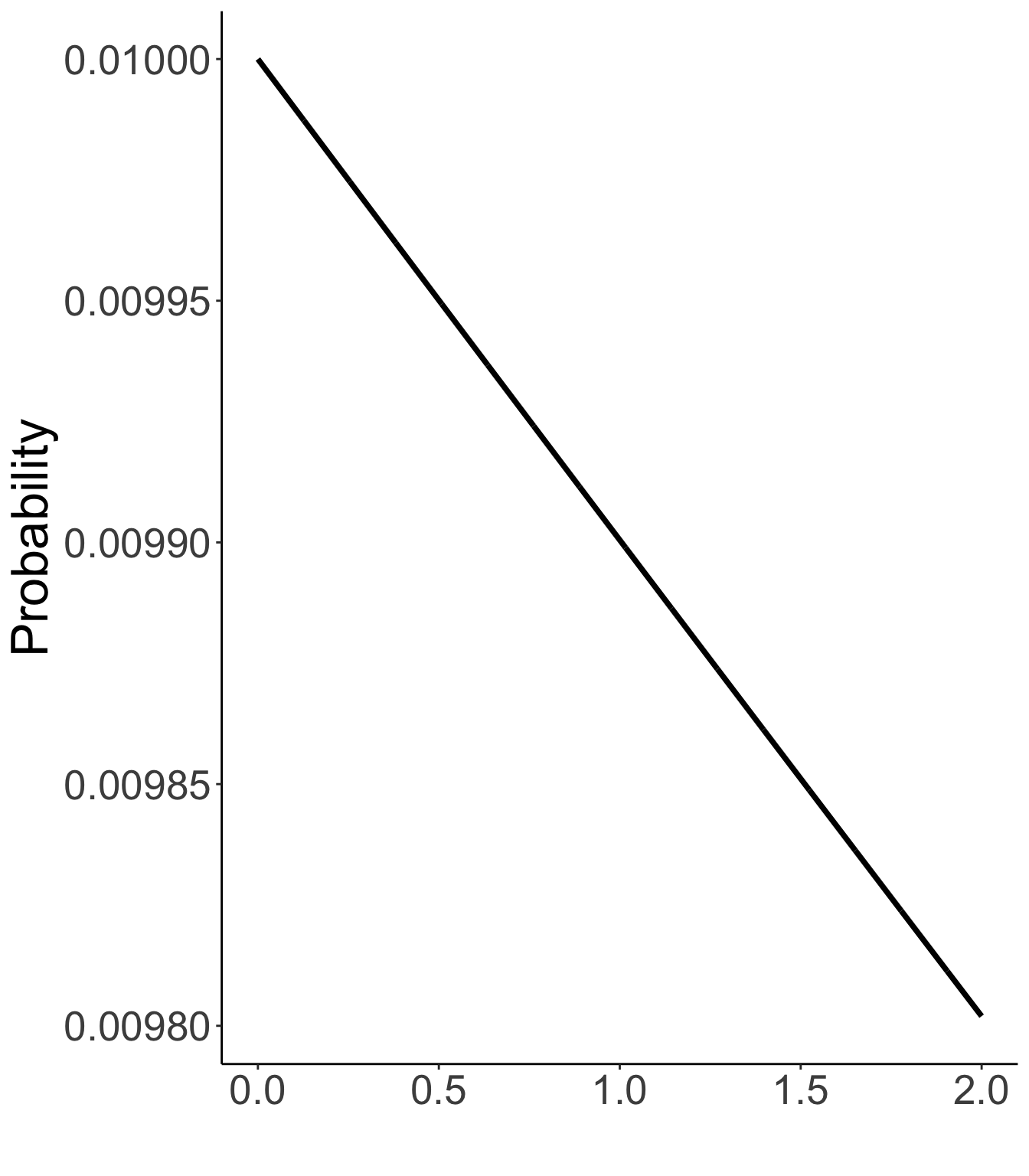
Prior distribution of unique standard deviations
Exp(.01)
```{r}
#| eval: true
#| echo: false
#| code-fold: true
#| fig-height: 8
data.frame(
x = seq(0, 2, .001),
y = dexp(x = seq(0, 2, .001), rate = .01)
) |>
ggplot(aes(x=x, y =y)) +
geom_path(linewidth = 1.3) +
labs(x = "", y = "Probability") +
theme_classic() +
theme(text = element_text(size = 25))
```
:::
:::::
## R's Data List Object
```{r}
#| eval: false
nObs = nrow(conspiracyItems)
nItems = ncol(conspiracyItems)
# item intercept hyperparameters
muMeanHyperParameter = 0
muMeanVecHP = rep(muMeanHyperParameter, nItems)
muVarianceHyperParameter = 1000
muCovarianceMatrixHP = diag(x = sqrt(muVarianceHyperParameter), nrow = nItems)
# item discrimination/factor loading hyperparameters
lambdaMeanHyperParameter = 0
lambdaMeanVecHP = rep(lambdaMeanHyperParameter, nItems)
lambdaVarianceHyperParameter = 1000
lambdaCovarianceMatrixHP = diag(x = sqrt(lambdaVarianceHyperParameter), nrow = nItems)
# unique standard deviation hyperparameters
psiRateHyperParameter = .01
psiRateVecHP = rep(.1, nItems)
modelCFA_data = list(
nObs = nObs,
nItems = nItems,
Y = conspiracyItems,
meanMu = muMeanVecHP,
covMu = muCovarianceMatrixHP,
meanLambda = lambdaMeanVecHP,
covLambda = lambdaCovarianceMatrixHP,
psiRate = psiRateVecHP
)
```
Note: In Stan, the second argument to the “normal” function is the **standard deviation** (i.e., the scale), not the variance (as in Bayesian Data Analysis) and not the inverse-variance (i.e., precision) (as in BUGS).
## Running the model in Stan
The total number of parameters is <mark>207</mark>.
- 177 person parameters ($\theta_1$ to $\theta_{177}$)
- 10 estimated parameters for item intercepts ($\mu_{1-10}$), factor loadings ($\lambda_{1-10}$), and unique standard deviation ($\psi_{1-10}$).
```{r}
#| eval: true
#| echo: false
root_dir <- "teaching/2024-01-12-syllabus-adv-multivariate-esrm-6553/Lecture07/Code"
save_dir <- "~/Library/CloudStorage/OneDrive-Personal/2024_Spring/ESRM6553 - Advanced Multivariate Modeling/Lecture07"
modelCFA_samples <- readRDS(here(save_dir, "model01.RDS"))
```
```{r}
#| eval: true
modelCFA_samples$metadata()$model_params
```
## Running the model in Stan
Different seed will have different initial values that may leads to convergence.
- `cmdstanr` sampling call:
```{r}
#| eval: false
modelCFA_samples = modelCFA_stan$sample(
data = modelCFA_data,
seed = 09102022,
chains = 4,
parallel_chains = 4,
iter_warmup = 1000,
iter_sampling = 2000
)
```
- The running time is about 2 seconds on my Macbook M1Pro
```{r}
#| eval: true
#| echo: false
modelCFA_samples$time()
```
- Note: Typically, longer chains are needed for larger models like this
- These will become even more longer when we use non-normal distributions for observed data
## Model Results
- Checking convergence with $\hat R$ (PSRF):
- The maximum of $\hat R$ is
```{r}
#| eval: true
#| echo: false
max(modelCFA_samples$summary(c("mu", "lambda", "psi"))[["rhat"]])
```
- Item Response Results
```{r}
#| eval: true
#| echo: false
#| html-table-processing: none
library(kableExtra)
kable(modelCFA_samples$summary(c("mu", "lambda", "psi")), digits = 2, simple = "pipe") |>
kable_classic(c("hover"), full_width = F)
```
## Model Results: Relationship between item means and mu parameters
```{r}
#| eval: true
#| echo: false
kable(cbind(ObservedMean = colMeans(conspiracyItems, na.rm = TRUE), modelCFA_samples$summary("mu")), digits = 3) |>
kable_classic(c("hover"), full_width = F)
```
<mark>Q1: The U.S. invasion of Iraq was not part of a campaign to fight terrorism, but was driven by oil companies and Jews in the U.S. and Israel.</mark>
Q1 has the highest agreement level for those with average conspiracy belief
## Modeling Strategy vs. Didactic Strategy
At this point, one should investigate model fit of the model
- If the model does not fit, then all model parameters could be biased
- Both item parameters ($\mu_i$, $\psi_i$ ) and person parameters ($\theta_p$)
- Moreover, the uncertainty accompanying each parameter (the posterior standard deviation) also be biased
- Especially bad for psychometric models as we quantify reliability with these number
But, to teach generalized measurement models, we will talk about differing models for observed data
- Different distributions
- Different parametrizations across different distributions
## Investigating Item Parameter
One plot that can help provide information about the item parameters is the **item characteristic curve** (ICC)
- Not called this in Confirmatory Factor Analysis (but equivalent)
- The ICC is the plot of the expected value of the response conditional on the value of the latent traits, for a range of latent trait values
$$
E(Y_{pi}\mid\theta_p)=\mu_i+\lambda_i\theta_p
$$
$$
ICC = f(E(Y_{pi}\mid\theta_p), \theta_p)
$$
- Because we have sampled values for each parameter, we can plot one ICC for each posterior draw
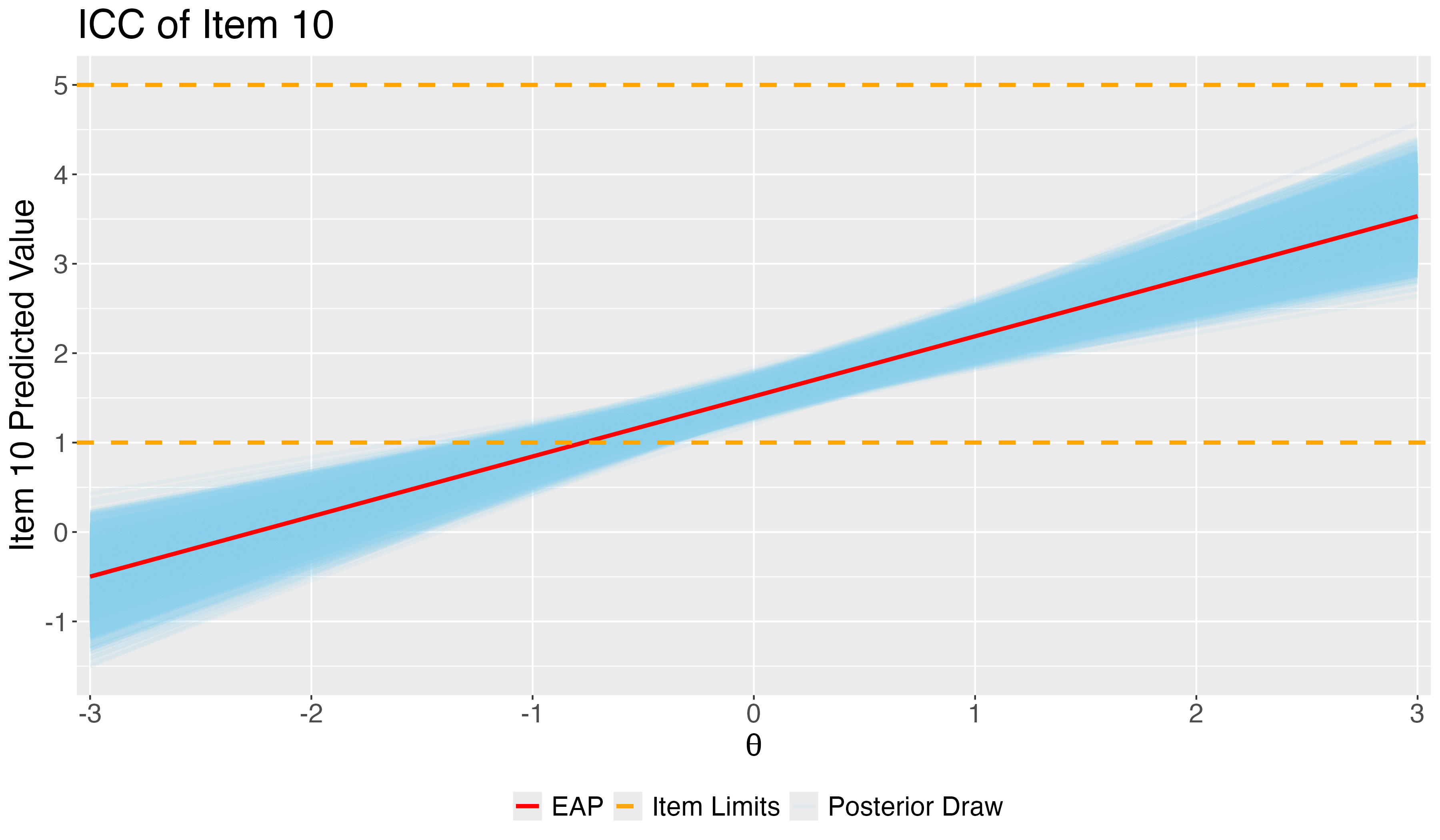
## Posterior ICC Plots: Difficulty and Discrimination of Item 10
- Using ICC, we can check the predicted response on item 10 given one's belief in conspiracy.
## Review Our R files
## Next Class
- More results from conspiracy data
- Compare results to `blavaan` package