---
title: "Lecture 04: ANOVA Assumptions Checking"
subtitle: "Experimental Design in Education"
date: "2025-02-10"
execute:
eval: true
echo: true
format:
html:
code-tools: true
code-line-numbers: false
code-fold: true
code-summary: 'Click to see R code'
number-offset: 1
fig.width: 10
fig-align: center
message: false
grid:
sidebar-width: 350px
uark-revealjs:
chalkboard: true
embed-resources: false
code-fold: false
number-sections: true
number-depth: 1
footer: "ESRM 64503"
slide-number: c/t
tbl-colwidths: auto
scrollable: true
output-file: slides-index.html
mermaid:
theme: forest
engine: knitr
filters:
- webr
---
## Class Outline
- Go through three assumptions of ANOVA and their checking statistics
- Post-hoc tests for more group comparisons
- Example: Intervention and Verbal Acquisition
- After-class Exercise: Effect of Sleep Duration on Cognitive Performance
## Review ANOVA Procedure
1. **Set hypotheses**:
- **Null hypothesis** ($H_0$): All group means are equal.
- **Alternative hypothesis** ($H_A$): At least one group mean differs.
2. **Determine statistical parameters**:
- Significance level $\alpha$
- Degrees of freedom for between-group ($df_b$) and within-group ($df_w$)
- Find the critical F-value.
3. **Compute test statistic**:
- Calculate **F-ratio** based on between-group and within-group sum of squares.
4. **Compare results**:
- Either compare $F_{\text{obs}}$ with $F_{\text{crit}}$ or $p$-value with $\alpha$.
- If $p < \alpha$, reject $H_0$.
## ANOVA Type I Error
1. The way to compare the means of multiple groups (i.e., more than two groups)
- Compared to multiple t-tests, the ANOVA does not inflate Type I error rates.
- For F-test in ANOVA, Type I error rate is not inflated (testing the $H_0$ once a time)
- Type I error rate is inflated when we do multiple comparison (testing multiple $H_0$s given the same data)
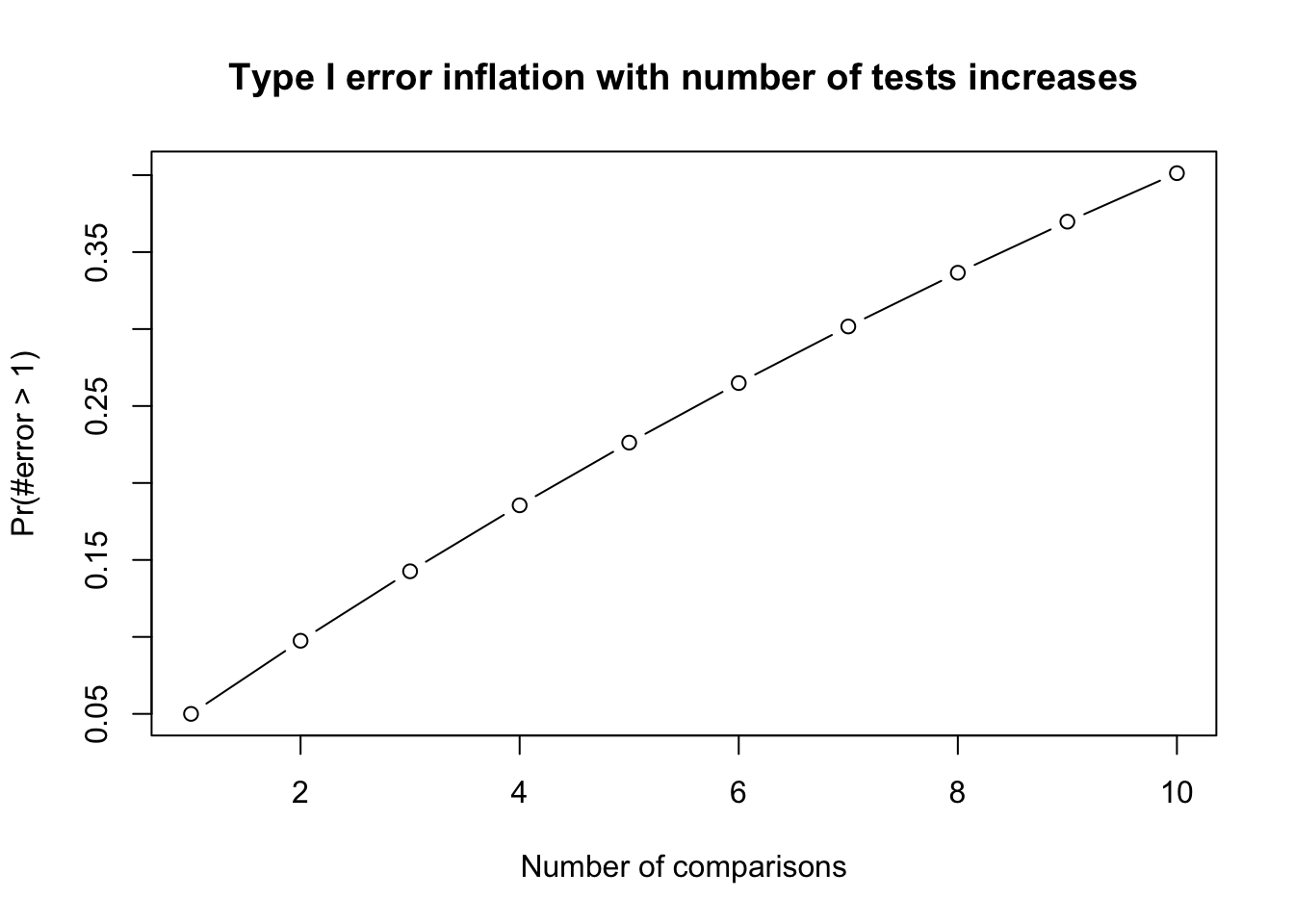
- **Family-wise error rate (FWER)**: probability of at least one Type I error occur = $1 − (1 − \alpha)^c$ where c =number of tests. Frequently used FWER methods include Bonferroni or FDR.
------------------------------------------
```{r}
#| code-fold: true
library(purrr)
alpha <- .05
c <- seq(10)
prob_TypeOneError <- 1 - (1 - alpha)^c
plot(x = c, y = prob_TypeOneError, type = "b",
xlab = "Number of comparisons", ylab = "Pr(#error > 1)",
main = "Type I error inflation with number of tests increases")
```
::: callout-note
## Family-Wise Error Rate
1. The **Family-Wise Error Rate** (FWER) is the probability of making at least one **Type I error** (false positive) across a set of multiple comparisons.
2. In **Analysis of Variance (ANOVA)**, multiple comparisons are often necessary when analyzing the differences between group means. FWER control is crucial to ensure the validity of statistical conclusions.
:::
# ANOVA Assumptions
## Overview
- Like all statistical tests, ANOVA requires certain assumptions to be met for valid conclusions:
- **Independence**: Observations are independent of each other.
- **Normality**: The residuals (errors) follow a normal distribution.
- **Homogeneity of variance (HOV)**: The variance within each group is approximately equal.
## Importance of Assumptions
- **If assumptions are violated**, the results of ANOVA may not be reliable.
- **We call models robust** when ANOVAs are not easily influenced by the violation of assumptions.
- Typically:
- ANOVA is **robust** to minor violations of normality, especially for large sample sizes (Central Limit Theorem).
- **Not robust** to violations of independence—if independence is violated, ANOVA is inappropriate.
- **Moderately robust** to HOV violations if sample sizes are equal.
## Assumption 1: Independence among samples
- **Definition**: Each observation should be independent of others.
- **Violations**:
- Clustering of data (e.g., repeated measures).
- Participants influencing each other (e.g., classroom discussions).
- **Check (Optional)**: Use the [Durbin-Watson test]{.redcolor} to check Serial Correlation.
- **Solution**: Repeated Measures ANOVA, Mixed ANOVA, Multilevel Model
- **Consequences**: If independence is violated, [ANOVA results are not valid]{.red}.
----------------------------------
**Example: Demonstrating Independence Violation**
```{r}
#| code-fold: true
#| results: hold
# Generate data with dependency (autocorrelation)
set.seed(1234)
n_per_group <- 20
groups <- 3
# Create dependent data using autoregressive structure
generate_dependent_data <- function(n, mean_val, sd_val, rho = 0.7) {
y <- numeric(n)
y[1] <- rnorm(1, mean_val, sd_val)
for(i in 2:n) {
y[i] <- rho * y[i-1] + rnorm(1, mean_val * (1-rho), sd_val * sqrt(1-rho^2))
}
return(y)
}
# Generate data for 3 groups with different means but with dependency
dependent_data <- data.frame(
group = factor(rep(c("A", "B", "C"), each = n_per_group)),
score = c(
generate_dependent_data(n_per_group, 50, 10), # Group A
generate_dependent_data(n_per_group, 55, 10), # Group B
generate_dependent_data(n_per_group, 60, 10) # Group C
)
)
# Compare with independent data
independent_data <- data.frame(
group = factor(rep(c("A", "B", "C"), each = n_per_group)),
score = c(
rnorm(n_per_group, 50, 10), # Group A
rnorm(n_per_group, 55, 10), # Group B
rnorm(n_per_group, 60, 10) # Group C
)
)
# Run ANOVA on both datasets
anova_dependent <- aov(score ~ group, data = dependent_data)
anova_independent <- aov(score ~ group, data = independent_data)
# Compare results
cat("ANOVA with DEPENDENT data:\n")
print(summary(anova_dependent))
cat("--------------------------\n")
cat("\nANOVA with INDEPENDENT data:\n")
print(summary(anova_independent))
# Check Durbin-Watson test
cat("--------------------------\n")
cat("\nDurbin-Watson test for dependent data:")
print(lmtest::dwtest(anova_dependent))
cat("--------------------------\n")
cat("\nDurbin-Watson test for independent data:")
print(lmtest::dwtest(anova_independent))
```
**Key observations:**
- Dependent data often shows artificially **smaller p-values** (inflated significance)
- Standard errors are **underestimated** when independence is violated
- Durbin-Watson test helps detect this violation (DW ≠ 2, p < 0.05)
------------------------------------------------------------------------
### Background: Durbin-Watson (DW) Test
::: callout-note
- The Durbin-Watson test is primarily used for detecting **autocorrelation** in time-series data.
- In the context of ANOVA with independent groups, residuals are generally assumed to be independent.
- It's good practice to check this assumption, especially if there's a reason to suspect potential autocorrelation.
:::
::: callout-important
- Properties of DW Statistic:
- $H_0$: [Independence of residual is satisfied.]{.redcolor}
- Ranges from 0 to 4.
- A value around 2 suggests no autocorrelation.
- Values approaching 0 indicate positive autocorrelation.
- Values toward 4 suggest negative autocorrelation.
- P-value
- P-Value: A small p-value (typically \< 0.05) indicates violation of independence
:::
---------------------------------------------
**Real-world examples of independence violations:**
- **Dependent data scenarios**:
- **Repeated measures**: Testing the same students multiple times (pre-test, mid-test, post-test)
- **Classroom clusters**: Students in the same classroom influence each other's performance
- **Time series**: Daily test scores where yesterday's performance affects today's
- **Spatial correlation**: Schools in the same district sharing similar resources and outcomes
- **Independent data scenarios**:
- **Random sampling**: Selecting students from different schools across different regions
- **Single measurement**: Each participant tested only once with no interaction
- **Controlled assignment**: Participants randomly assigned to groups with no cross-contamination
**Why this matters**: When observations are dependent, they don't provide as much unique information as independent observations. This leads to overconfident results (smaller standard errors and p-values).
---------------------------------------------
### Visualization of indepdence test
```{r}
#| code-fold: true
#| fig-width: 12
#| fig-height: 8
#| warning: false
library(ggplot2)
library(gridExtra)
# Use the same data from previous example
n_per_group <- 20
# Generate dependent and independent data
dependent_data <- data.frame(
group = factor(rep(c("A", "B", "C"), each = n_per_group)),
score = c(
generate_dependent_data(n_per_group, 50, 10),
generate_dependent_data(n_per_group, 55, 10),
generate_dependent_data(n_per_group, 60, 10)
),
observation = 1:(3*n_per_group),
type = "Dependent (AR)"
)
independent_data <- data.frame(
group = factor(rep(c("A", "B", "C"), each = n_per_group)),
score = c(
rnorm(n_per_group, 50, 10),
rnorm(n_per_group, 55, 10),
rnorm(n_per_group, 60, 10)
),
observation = 1:(3*n_per_group),
type = "Independent"
)
# Combine data
combined_data <- rbind(dependent_data, independent_data)
# Plot 1: Time series plot showing autocorrelation
p1 <- ggplot(combined_data, aes(x = observation, y = score, color = group)) +
geom_line(size = 0.8) + geom_point(size = 1.5) +
facet_wrap(~type, scales = "free") +
labs(title = "Time Series: Independence vs Dependence",
x = "Observation Order", y = "Score") +
theme_minimal() + theme(legend.position = "bottom")
# Plot 2: Lag plot to show autocorrelation
create_lag_data <- function(data) {
data$score_lag1 <- c(NA, data$score[-length(data$score)])
return(data[!is.na(data$score_lag1), ])
}
lag_dependent <- create_lag_data(dependent_data)
lag_independent <- create_lag_data(independent_data)
lag_combined <- rbind(lag_dependent, lag_independent)
p2 <- ggplot(lag_combined, aes(x = score_lag1, y = score)) +
geom_point(aes(color = group), alpha = 0.7) +
geom_smooth(method = "lm", se = TRUE, color = "black", size = 0.5) +
facet_wrap(~type, scales = "free") +
labs(title = "Lag Plot: Current vs Previous Observation",
x = "Score (t-1)", y = "Score (t)") +
theme_minimal() + theme(legend.position = "bottom")
# Display plots
grid.arrange(p1, p2, ncol = 1)
```
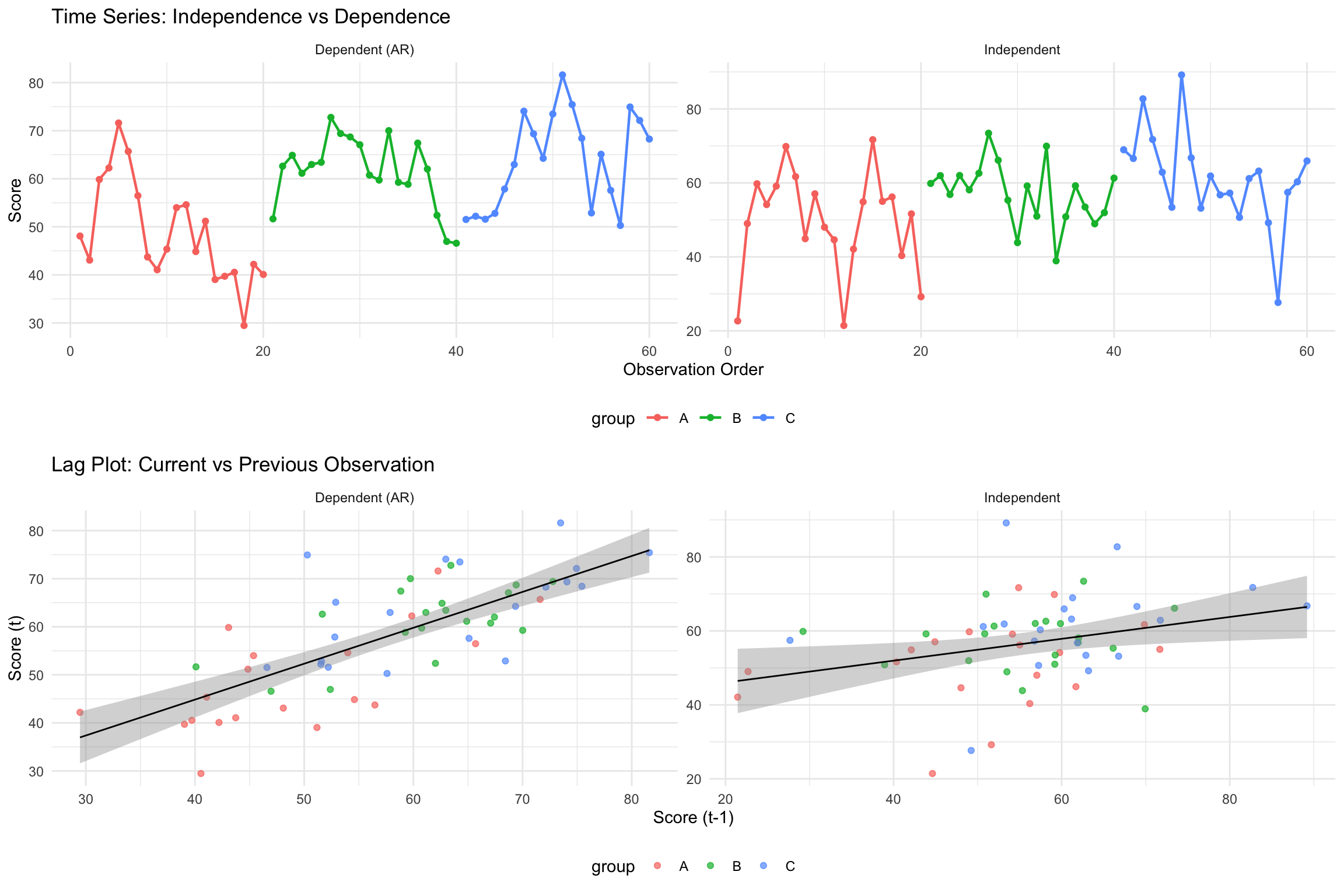
**Key Visual Differences:**
- **Dependent data**: Shows smooth, connected patterns with visible trends
- **Independent data**: Shows random scatter with no systematic patterns
- **Lag plot**: Dependent data shows positive correlation between consecutive observations
------------------------------------------------------------------------
[`lmtest::dwtest()`]{.bluecolor .bigger}
- Performs the Durbin-Watson test for autocorrelation of disturbances.
- The Durbin-Watson test has the null hypothesis that the autocorrelation of the disturbances is 0.
```{r}
#| eval: false
#install.packages("lmtest")
library(lmtest)
err1 <- rnorm(100)
## generate regressor and dependent variable
x <- rep(c(-1,1), 50)
y <- 1 + x + err1
dat <- data.frame(x = x, y = y)
## perform Durbin-Watson test
dwtest(y ~ x, data = dat)
```
These results suggest there is no autocorrelation at the alpha level of .05.
--------------------------------
::: panel-tabset
### Exercise
- Below is the R code that has loaded two datasets for you. `dat1` and `dat2`, try to examine which dataset violates independence.
```{webr-r}
#| context: setup
set.seed(1999)
# Create dependent data using autoregressive structure
generate_dependent_data <- function(n, mean_val, sd_val, rho = 0.7) {
y <- numeric(n)
y[1] <- rnorm(1, mean_val, sd_val)
for(i in 2:n) {
y[i] <- rho * y[i-1] + rnorm(1, mean_val * (1-rho), sd_val * sqrt(1-rho^2))
}
return(y)
}
# Use the same data from previous example
n_per_group <- 30
# Generate dependent and independent data
dat1 <- data.frame(
group = factor(rep(c("A", "B", "C"), each = n_per_group)),
score = c(
generate_dependent_data(n_per_group, 50, 10),
generate_dependent_data(n_per_group, 55, 10),
generate_dependent_data(n_per_group, 60, 10)
)
)
dat2 <- data.frame(
group = factor(rep(c("A", "B", "C"), each = n_per_group)),
score = c(
rnorm(n_per_group, 50, 10),
rnorm(n_per_group, 55, 10),
rnorm(n_per_group, 60, 10)
)
)
```
```{webr-r}
#| editor-code-line-numbers: 4-7
print(dat1)
print(dat2)
## Test which data violate independence assumption
library(________)
print(dwtest(________))
print(dwtest(________))
```
### Answer
```{webr-r}
# DW = 0.8248, p-value = 6.68e-11, dat1 violates independence
# DW = 2.2514, p-value = 0.8386, dat2 does not violates independence
library(lmtest)
print(dwtest(score ~ group, data = dat1))
print(dwtest(score ~ group, data = dat2))
```
:::
## Assumption 2: Normality
- The **dependent variable (DV)** should be normally distributed within each group.
- **Assessments**:
- Graphical methods: Histograms, [Q-Q plots]{.redcolor}.
- Statistical tests:
- [Shapiro-Wilk test (common)]{.redcolor}
- Kolmogorov-Smirnov (KS) test (for large samples)
- Anderson-Darling test (detects kurtosis issues).
- **Robustness**:
- ANOVA is robust to normality violations for large samples.
- If normality is violated, consider transformations or non-parametric tests.
------------------------------------------------------------------------
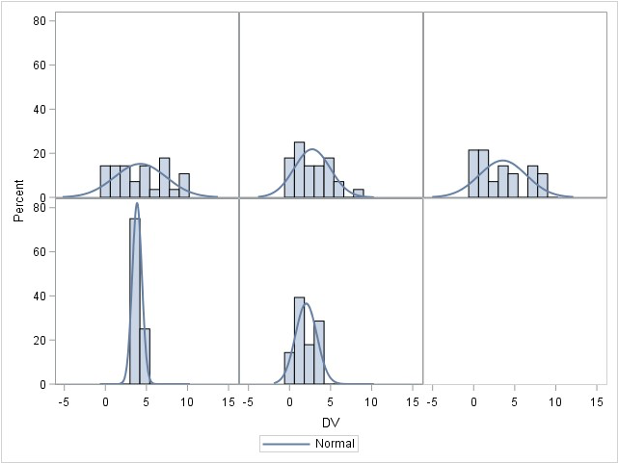
### Normality check for the whole samples
- Assume that the DV (Y) is distributed normally within each group for ANOVA
- ANOVA is **robust** to minor violations of normality
- So generally start with homogeneity, then assess normality
------------------------------------------------------------------------
### R function `shapiro.test()`
```{r}
#| eval: true
#| code-fold: true
#| results: hold
set.seed(123) # For reproducibility
sample_normal_data <- rnorm(200, mean = 50, sd = 10) # Generate normal data
sample_nonnormal_data <- sample(1:10, 200, replace = TRUE) # Generate non-normal data
cat("Normality assumption holds:")
shapiro.test(sample_normal_data)
cat("Normality assumption does not hold:")
shapiro.test(sample_nonnormal_data)
```
------------------------------------------------------------------------
### R function `ks.test()`
```{r}
#| results: hold
# Perform Kolmogorov-Smirnov test against a normal distribution
cat("Normality assumption holds:")
ks.test(scale(sample_normal_data), "pnorm")
cat("Normality assumption does not hold:")
ks.test(scale(sample_nonnormal_data), "pnorm")
```
::: panel-tabset
### Exercise II
Perform Shapiro-Wilk and Kolmogorov-Smirnov for both datasets. Tell me which test works or does not work and why.
```{webr-r}
#| editor-code-line-numbers: 6-11
set.seed(2345)
normal_data <- rnorm(50, mean = 50, sd = 10) # normal data
nonnormal_data <- sample(1:10, 50, replace = TRUE) # non-normal
### Perform Shapiro-Wilk and Kolmogorov-Smirnov for both data
### Shapiro-Wilk
_____(normal_data)
_____(nonnormal_data)
### Kolmogorov-Smirnov
_____(____(normal_data, _____))
_____(____(nonnormal_data, _____))
```
### Answer
```{webr-r}
shapiro.test(normal_data)
shapiro.test(nonnormal_data)
ks.test(scale(normal_data), "pnorm")
ks.test(scale(nonnormal_data), "pnorm")
```
:::
-----------------------
### Normaliy check for each group
```{r}
#| code-fold: false
#| warning: false
# Generate three-group data with normal distributions
library(dplyr) # we need this package to group the data
set.seed(123)
group_data <- data.frame(
group = rep(c("A", "B", "C"), each = 30),
score = c(rnorm(30, mean = 50, sd = 10), # Group A
rnorm(30, mean = 55, sd = 12), # Group B
rnorm(30, mean = 48, sd = 9)) # Group C
)
# Run Shapiro-Wilk test for each group
group_data |>
group_by(group) |>
summarize(
shapiro_p_value = shapiro.test(score)$p.value
)
```
**Interpretation of Normality Checking Results:**
- If all p-values > 0.05: All groups satisfy normality assumption - proceed with ANOVA
- If one group has p-value < 0.05 but others are normal:
- ANOVA is generally robust to minor normality violations, especially with equal sample sizes
- Consider the sample size: larger samples (n > 30) make ANOVA more robust
- Options: (1) Continue with ANOVA if violation is mild, (2) Apply data transformation (log, square root), or (3) Use non-parametric alternative (Kruskal-Wallis test)
- If multiple groups violate normality: Strong evidence against using ANOVA - consider transformations or non-parametric tests
--------
::: panel-tabset
### Exercise III
Perform Shapiro-Wilk for each group datasets. Tell me which group is not normal.
```{webr-r}
#| context: setup
set.seed(2345)
group_data2 <- data.frame(
group = rep(c("A", "B", "C"), each = 30),
score = c(round(rnorm(30, mean = 50, sd = 10), 2), # Group A
sample(seq(50, 100, .01), 30, replace = TRUE), # Group B
round(rnorm(30, mean = 48, sd = 9), 2)) # Group C
)
```
```{webr-r}
#| editor-code-line-numbers: 5-9
library(dplyr) # we need this package to group the data
head(group_data2) # check first 6 rows
### Perform Shapiro-Wilk for each group
group_data2 |>
______(______) |>
______(
shapiro_p_value = __________(________)$__________
)
```
### Answer
```{webr-r}
group_data2 |>
group_by(group) |>
summarize(
shapiro_p_value = shapiro.test(score)$p.value
)
```
:::
--------------------------------------------------------------------
## Assumption 3: Homogeneity of Variance (HOV)
- Variance across groups should be equal.
- **Assessments**:
- **Levene’s test**: Tests equality of variances.
- **Brown-Forsythe test**: More robust to non-normality.
- **Bartlett's test**: For data with normality.
- **Boxplots**: Visual inspection.
- **What if violated?**
- **Welch’s ANOVA** (adjusted for variance differences).
- **Transforming** the dependent variable.
- **Using non-parametric tests** (e.g., Kruskal-Wallis).
------------------------------------------------------------------------
[Practical Considerations]{.bluecolor .bigger}
```{r}
#| eval: false
## Computes Levene's test for homogeneity of variance across groups.
car::leveneTest(outcome ~ group, data = data)
## Boxplots to visualize the variance by groups
boxplot(outcome ~ group, data = data)
## Brown-Forsythe test
onewaytests::bf.test(outcome ~ group, data = data)
## Bartlett's Test
bartlett.test(outcome ~ group, data = data)
```
- **Levene's test** and **Brown-Forsythe test** are often preferred when data does not meet the assumption of normality, especially for small sample sizes.
- **Bartlett’s test** is most powerful when the data is normally distributed and when sample sizes are equal across groups. However, it can be less reliable if these assumptions are violated.
------------------------------------------------------------------------
[Decision Tree]{.redcolor .bigger}
```{dot}
//| echo: false
digraph flowchart {
node [shape=box, style=filled, fillcolor=lightgray];
A [label="Is the data normal by group?"];
B [label="Use Bartlett's Test"];
C [label="Are there outliers?"];
D [label="Use Brown-Forsythe Test"];
E [label="Use Levene's Test"];
A -> B [label="Yes"];
A -> C [label="No"];
C -> D [label="Yes"];
C -> E [label="No"];
}
```
------------------------------------------------------------------------
## ANOVA Robustness
- **Robust to**:
- Minor normality violations (for large samples).
- Small HOV violations if group sizes are **equal**.
- **Not robust to**:
- **Independence violations**—ANOVA is invalid if data points are dependent.
- **Severe HOV violations**—Type I error rates become unreliable.
- The robustness of assumptions is something you should be careful about before/when you perform data collection. They are not something you can address after data collection has been finished.
------------------------------------------------------------------------
[Robustness to Violations of Normality Assumption]{.redcolor}
- ANOVA assumes that the residuals (errors) are normally distributed within each group.
- However, ANOVA is generally robust to violations of normality, particularly when **the sample size is large**.
- **Theoretical Justification**: This robustness is primarily due to the Central Limit Theorem (CLT), which states that, for sufficiently large sample sizes (typically $n≥30$ per group), the sampling distribution of the mean approaches normality, even if the underlying population distribution is non-normal.
- This means that unless the data are heavily skewed or have extreme outliers, ANOVA results remain valid and Type I error rates are not severely inflated.
------------------------------------------------------------------------
### Robustness to Violations of Homogeneity of Variance
- The homogeneity of variance (homoscedasticity) assumption states that all groups should have equal variances. ANOVA can tolerate moderate violations of this assumption, particularly when:
- **Sample sizes are equal (or nearly equal) across groups** – When groups have equal sample sizes, the F-test remains robust to variance heterogeneity because the pooled variance estimate remains balanced.
- **The degree of variance heterogeneity is not extreme** – If the largest group variance is no more than about four times the smallest variance, ANOVA results tend to remain accurate.
------------------------------------------------------------------------
### ANOVA: Lack of Robustness to Violations of Independence of Errors
- The assumption of independence of errors means that observations within and between groups must be uncorrelated. Violations of this assumption severely compromise ANOVA’s validity because:
- **Inflated Type I error rates** – If errors are correlated (e.g., due to clustering or repeated measures), standard errors are underestimated, leading to an increased likelihood of falsely rejecting the null hypothesis.
- **Biased parameter estimates** – When observations are not independent, the variance estimates do not accurately reflect the true variability in the data, distorting F-statistics and p-values.
- **Common sources of dependency** – Examples include nested data (e.g., students within schools), repeated measurements on the same subjects, or time-series data. In such cases, alternatives like mixed-effects models or generalized estimating equations (GEE) should be considered.
# Omnibus ANOVA Test
## Overview
- **What does it test?**
- Whether there is **at least one** significant difference among means.
- **Limitation**:
- Does **not** tell **which** groups are different.
- **Solution**:
- Conduct **post-hoc tests**.
## Individual Comparisons of Means
- If ANOVA is **significant**, follow-up tests identify **where** differences occur.
- **Types**:
- **Unplanned (post-hoc) comparisons**: Conducted **after** ANOVA (Ominibus ANOVA Test; Post-hoc Test).
- **Planned comparisons**: Defined **before** data collection.
## Planned vs. Unplanned Comparisons
- **Unplanned (post-hoc)**:
- **Data-driven**.
- Only performed **if ANOVA is significant**.
- **Planned**:
- Based on **theory**.
- Can be done **even if ANOVA is not significant**.
## Types of Unplanned Comparisons
- **Common post-hoc tests**:
1. **Tukey’s HSD**
2. **Family-Wise Error Rate** or Adjusted p-values
3. **Fisher’s LSD**
4. **Sidák correction**
------------------------------------------------------------------------
### Post-hoc test
[Tukey’s HSD]{.bluecolor}
- Controls for **Type I error** across multiple comparisons.
- Uses a **q-statistic** from a Tukey table.
- Preferred when **all pairs** need comparison.
```{r}
aov_result <- aov(score ~ group, group_data) # run the ANOVA model first
TukeyHSD(aov_result) # call TukeyHSD followed by the model to perform Tukey multiple comparisons
```
[Family-wise Error Rate (adjusted p-values)]{.bluecolor}
- Adjusts p-values to reduce Type I error
- Report adjusted p-values (typically larger that original p-values)
### Tukey Honest Significant Differences (HSD) Test
- **Purpose**: Conducts all possible pairwise comparisons between group means while controlling the family-wise error rate at the specified alpha level (typically 0.05).
- **Why it's needed**: Simple pairwise t-tests after a significant ANOVA inflate the Type I error rate. Tukey HSD adjusts for multiple comparisons to maintain the overall error rate.
- **How it works**: Creates confidence intervals for all pairwise mean differences using the studentized range distribution, which accounts for the number of groups being compared.
- **Interpretation**: If a confidence interval does not include zero, or if p adj < 0.05, the pair of groups differs significantly.
```{r}
#| eval: true
tukey_result <- TukeyHSD(aov_result)
print(tukey_result$group)
```
**Interpretation**: The Tukey HSD test provides pairwise comparisons between all groups while controlling for family-wise error rate. The output shows:
- **diff**: The difference in means between each pair of groups
- **lwr & upr**: The 95% confidence interval for the difference
- **p adj**: The adjusted p-value for multiple comparisons
Pairs with p adj < 0.05 indicate statistically significant differences between those groups.
------------------------------------------------------------------------
```{r}
#| eval: true
plot(tukey_result)
```
## Interpreting Results
### ANOVA Statistics
```
Df Sum Sq Mean Sq F value Pr(>F)
group 3 724.1 241.37 11.45 2.04e-05 ***
Residuals 36 759.2 21.09
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
```
```{r}
#| eval: false
#| code-fold: true
library(dplyr)
data |>
group_by(group) |>
summarise(
Mean = mean(score),
SD = sd(score)
)
```
- A one-way analysis of variance (ANOVA) was conducted to examine the effect of teaching method on students' test scores. The results indicated a statistically significant difference in test scores across the four groups, (F(3, 36) = 11.45, p < .001). Post-hoc comparisons using the Tukey HSD test revealed that the Interactive method (G3) (M = 28.06, SD = 3.33) resulted in significantly higher scores than the Traditional method (M = 18.17, SD = 4.47) with p < .001. However, no significant difference was found between G1 and the Control group (p = .99). These results suggest that certain teaching methods can significantly improve student performance compared to traditional approaches.
## ANOVA Example: Intervention and Verbal Acquisition
### Background
- Research Question: Does an intensive intervention improve students’ verbal acquisition scores?
- Study Design:
- 4 groups: Control, G1, G2, G3 (treatment levels).
- Outcome variable: Verbal acquisition score (average of three assessments).
- Hypotheses:
- $H_0$: No difference in verbal acquisition scores across groups.
- $H_A$: At least one group has a significantly different mean.
### Step 1: Generate Simulated Data in R
```{r}
#| warning: false
#| message: false
#| results: hold
# Load necessary libraries
library(tidyverse)
# Set seed for reproducibility
set.seed(123)
# Generate synthetic data for 4 groups
data <- tibble(
group = rep(c("Control", "G1", "G2", "G3"), each = 30),
verbal_score = c(
rnorm(30, mean = 70, sd = 10), # Control group
rnorm(30, mean = 75, sd = 12), # G1
rnorm(30, mean = 80, sd = 10), # G2
rnorm(30, mean = 85, sd = 8) # G3
)
)
# View first few rows
head(data)
```
**Function explanations:**
- `library(tidyverse)`: Loads the tidyverse package collection, including dplyr, ggplot2, and tibble
- `set.seed(123)`: Sets the random number generator seed to ensure reproducible results across runs
- `tibble()`: Creates a modern data frame (tibble) with improved printing and subsetting behavior
- `rep(c("Control", "G1", "G2", "G3"), each = 30)`: Repeats each group label 30 times, creating 120 total observations (30 per group)
- `rnorm(30, mean = X, sd = Y)`: Generates 30 random numbers from a normal distribution with specified mean and standard deviation
- `c()`: Combines the four vectors of random numbers into a single vector of 120 values
- `head(data)`: Displays the first 6 rows of the dataset for preview
### Step 2: Summary Statistics
```{r}
# Summary statistics by group
data|>
group_by(group)|>
summarise(
mean_score = mean(verbal_score),
sd_score = sd(verbal_score),
n = n()
)
```
**Function explanations:**
- `group_by(group)`: Groups the data by the `group` variable, allowing subsequent operations to be performed separately for each group
- `summarise()`: Creates summary statistics for each group, collapsing the grouped data into summary values
- `mean(verbal_score)`: Calculates the arithmetic mean of verbal scores within each group
- `sd(verbal_score)`: Computes the standard deviation of verbal scores within each group
- `n()`: Counts the number of observations in each group
------------------------------------------------------------------------
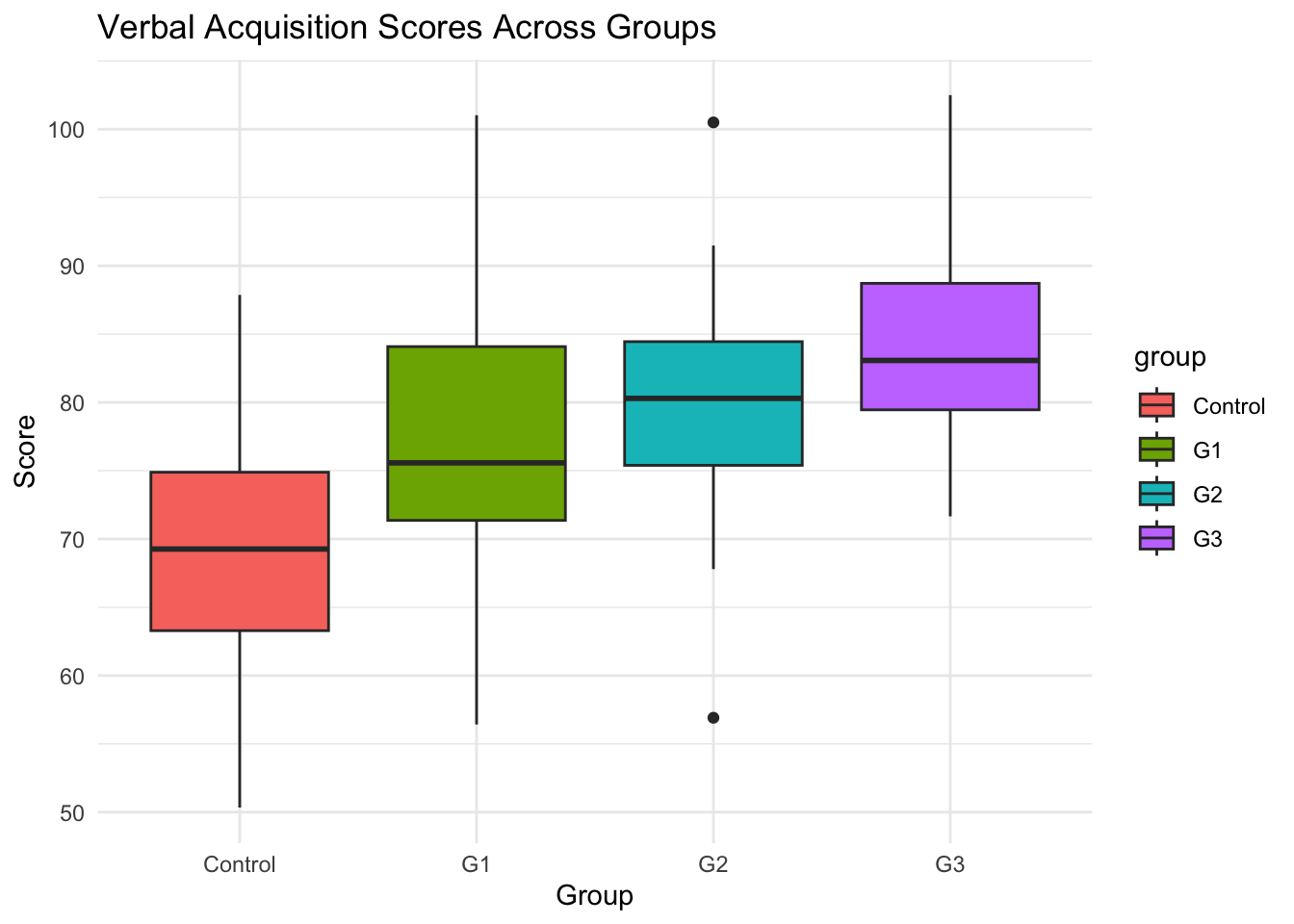
```{r}
# Boxplot visualization
library(ggplot2) # Load ggplot2 package for creating visualizations
ggplot(data, aes(x = group, y = verbal_score, fill = group)) + # Create base plot with group on x-axis, scores on y-axis, fill by group
geom_boxplot() + # Add boxplot layer to show distribution of scores within each group
theme_minimal() + # Apply minimal theme for clean appearance
labs(title = "Verbal Acquisition Scores Across Groups", y = "Score", x = "Group") # Add descriptive labels
```
### Step 3: Check ANOVA Assumptions
### Assumption Check 1: Independence of residuals Check
```{r}
#| results: hold
#| warning: false
#| message: false
# Fit the ANOVA model
anova_model <- lm(verbal_score ~ group, data = data)
# Load the lmtest package
library(lmtest)
# Perform the Durbin-Watson test
dw_test_result <- dwtest(anova_model)
# View the test results
print(dw_test_result)
```
- Interpretation:
- In this example, the `DW` value is close to 2, and the p-value is greater than 0.05, indicating no significant autocorrelation in the residuals.
------------------------------------------------------------------------
### Assumption Check 2: Normality Check
```{r}
# Shapiro-Wilk normality test for each group
data |>
group_by(group) |>
summarise(
shapiro_p = shapiro.test(verbal_score)$p.value
)
```
- Interpretation:
- If $p>0.05$, normality assumption is not violated.
- If $p<0.05$, data deviates from normal distribution.
- Since the data meets the normality requirement and no outliers, we can use [Bartlett's or Levene's tests]{.redcolor} for HOV checking.
------------------------------------------------------------------------
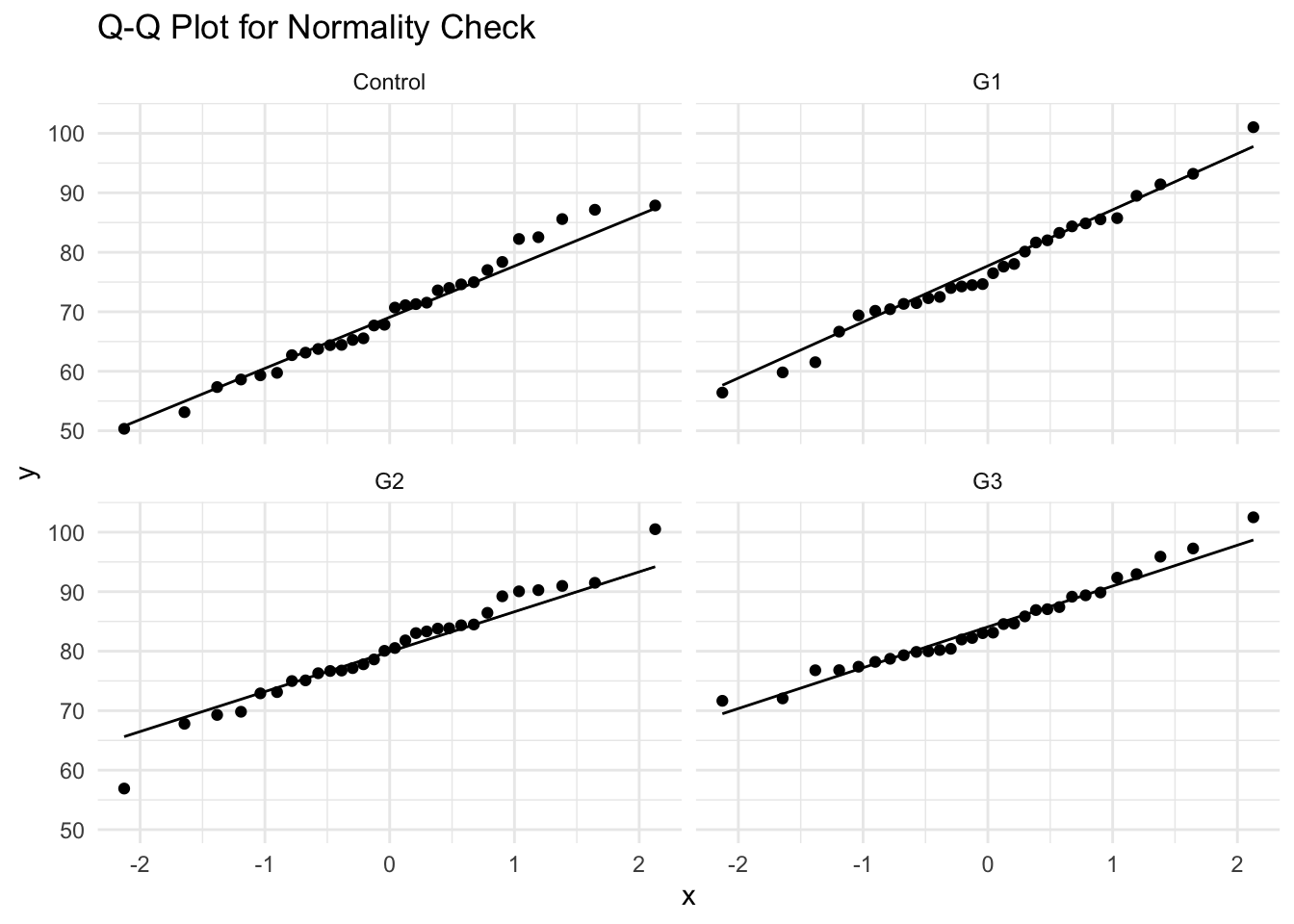
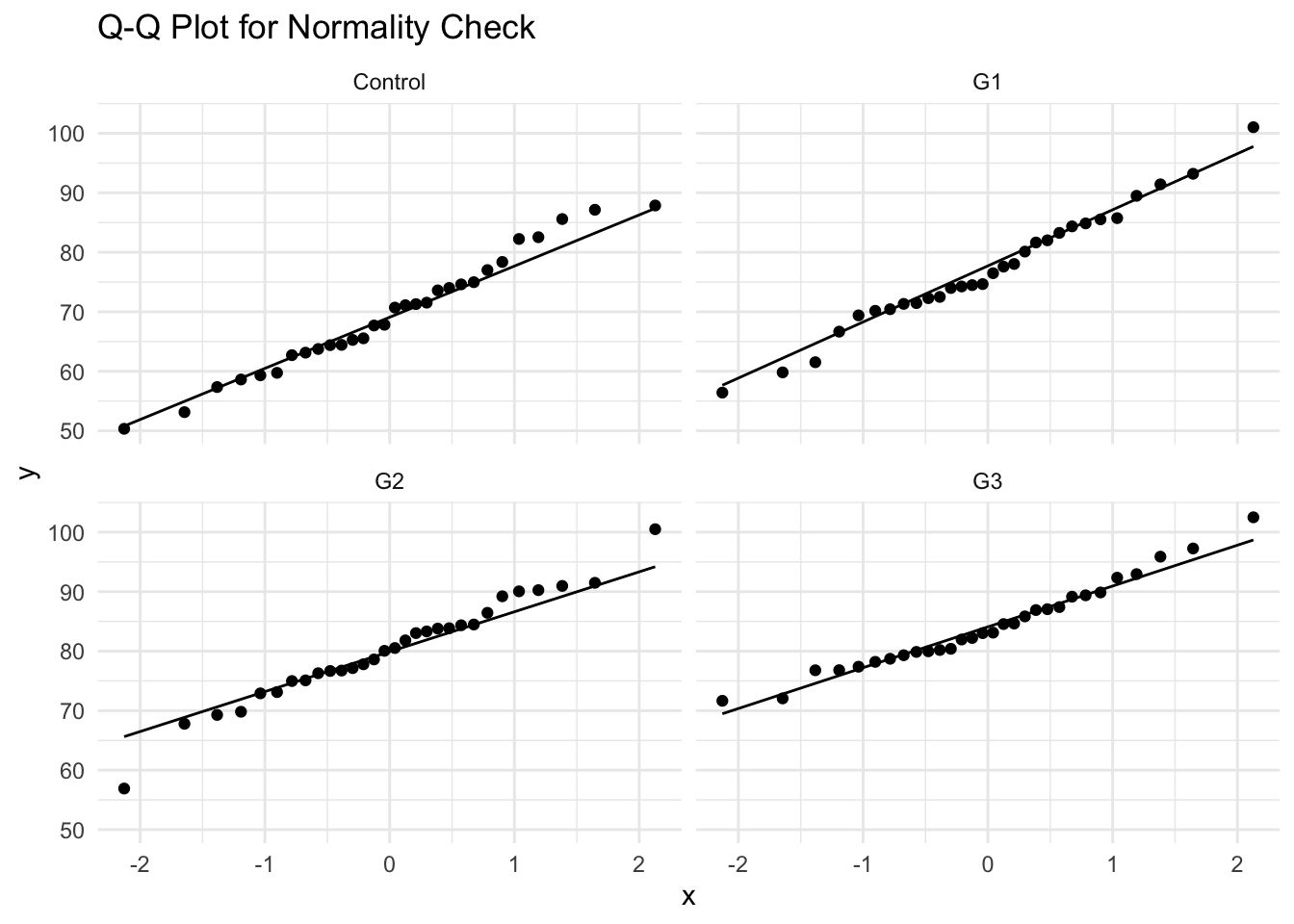
- Alternative Check: Q-Q Plot
```{r}
ggplot(data, aes(sample = verbal_score)) +
geom_qq() + geom_qq_line() +
facet_wrap(~group) +
theme_minimal() +
labs(title = "Q-Q Plot for Normality Check")
```
------------------------------------------------------------------------
### Assumption Check 3: Homogeneity of Variance (HOV) Check
```{r}
#| results: hold
#| warning: false
library(car) # for leveneTest
library(onewaytests) # for bf.test
data$group <- as.factor(data$group)
car::leveneTest(verbal_score ~ group, data = data)
cat("-----------------------------\n")
bartlett.test(verbal_score ~ group, data = data)
cat("-----------------------------\n")
onewaytests::bf.test(verbal_score ~ group, data = data)
```
- Interpretation:
- If $p>0.05$, variance is homogeneous (ANOVA assumption met).
- If $p<0.05$, variance differs across groups (consider Welch’s ANOVA).
- It turns out our data does not violate the homogeneity of variance assumption.
### Step 4: Perform One-Way ANOVA
```{r}
anova_model <- aov(verbal_score ~ group, data = data)
summary(anova_model)
```
- Interpretation:
- If $p<0.05$, at least one group mean is significantly different.
- If $p>0.05$, fail to reject $H0$ (no significant differences).
### Step 5: Post-Hoc Tests (Tukey's HSD)
- After finding a significant ANOVA result, we perform Tukey's HSD test to identify which specific groups differ from each other.
- For example, we can compare:
- G1 vs Control
- G2 vs Control
- G3 vs Control
- G2 vs G1
- G3 vs G1
- G3 vs G2
```{r}
#| code-fold: false
# Tukey HSD post-hoc test
tukey_results <- TukeyHSD(anova_model)
round(tukey_results$group, 3)
```
- Interpretation:
- Identifies which groups differ.
- If $p<0.05$, the groups significantly differ.
- G1-Control
- G2-Control
- G3-Control
- G3-G1
------------------------------------------------------------------------
### Step 6: Family-wise Error Rate Control
- General Linear Hypothesis Tests: `glht` function in multcomp package allow us to perform family-wise error rate control (using `adjusted p-values`).
- Alternatively, we can use `TukeyHSD` to adjust p values to control inflated Type I error.
$$
Y = \beta_0G_0+\beta_1G_1 +\beta_2G_2 +\beta_3G_3
$$
- When $\{G_0, G_1, G_2, G_3\}$ = $\{0, 1, 0, 0\}$, $Y = \beta_1$, which is the group difference between G1 with the control group
- When $\{G_0, G_1, G_2, G_3\}$ = $\{0, 0, 1, 0\}$, $Y = \beta_2$, which is the group difference between G2 with the control group
- When $\{G_0, G_1, G_2, G_3\}$ = $\{0, -1, 1, 0\}$, $Y = \beta_2 - \beta_1$, which is the group difference between G2 with G1
```{r}
#| warning: false
#| message: false
#| results: hold
library(multcomp)
# install.packages("multcomp")
### set up multiple comparisons object for all-pair comparisons
# head(model.matrix(anova_model))
comprs <- rbind(
"G1 - Ctrl" = c(0, 1, 0, 0),
"G2 - Ctrl" = c(0, 0, 1, 0),
"G3 - Ctrl" = c(0, 0, 0, 1),
"G2 - G1" = c(0, -1, 1, 0),
"G3 - G1" = c(0, -1, 0, 1),
"G3 - G2" = c(0, 0, -1, 1)
)
cht <- glht(anova_model, linfct = comprs)
summary(cht, test = adjusted("fdr"))
```
------------------------------------------------------------------------
### Side-by-Side comparison of two methods
::::: columns
::: {.column width="50%"}
#### TukeyHSD method
```{r}
#| eval: false
#| code-line-numbers: false
#| code-summary: "TukeyHSD method"
diff lwr upr p adj
G1-Control 7.611 1.545 13.677 0.008 **
G2-Control 10.715 4.649 16.782 0.000 ***
G3-Control 14.720 8.654 20.786 0.000 ***
G2-G1 3.104 -2.962 9.170 0.543
G3-G1 7.109 1.042 13.175 0.015 *
G3-G2 4.005 -2.062 10.071 0.318
```
:::
::: {.column width="50%"}
#### FDR method
```{r}
#| eval: false
#| code-line-numbers: false
#| code-summary: "multcomp package"
Estimate Std. Error t value Pr(>|t|)
G1 - Ctrl == 0 7.611 2.327 3.270 0.00283 **
G2 - Ctrl == 0 10.715 2.327 4.604 3.20e-05 ***
G3 - Ctrl == 0 14.720 2.327 6.325 2.94e-08 ***
G2 - G1 == 0 3.104 2.327 1.334 0.18487
G3 - G1 == 0 7.109 2.327 3.055 0.00419 **
G3 - G2 == 0 4.005 2.327 1.721 0.10555
```
:::
- The differences in p-value adjustment between the `TukeyHSD` method and the `multcomp` package stem from how each approach calculates and applies the multiple comparisons correction. Below is a detailed explanation of these differences.
:::::
### Comparison of Differences
| Feature | `TukeyHSD()` (Base R) | `multcomp::glht()` |
|----|----|----|
| **Distribution Used** | Studentized Range (q-distribution) | t-distribution |
| **Error Rate Control** | Strong FWER control | Flexible error control |
| **Simultaneous Confidence Intervals** | Yes | Typically not (depends on method used) |
| **Adjustment Method** | Tukey-Kramer adjustment | Single-step, Westfall, Holm, Bonferroni, etc. |
| **P-value Differences** | More conservative (larger p-values) | Slightly different due to t-distribution |
### Step 6: Reporting ANOVA Results
[Result]{.bigger}
1. We first examined three assumptions of ANOVA for our data as the preliminary analysis. According to the Durbin-Watson test, the Shapiro-Wilk normality test, and the Bartletts' test, the sample data meets all assumptions of the one-way ANOVA modeling.
2. A one-way ANOVA was then conducted to examine the effect of three intensive intervention methods (Control, G1, G2, G3) on verbal acquisition scores. There was a statistically significant difference between groups, $F(3,116)=14.33$, $p<.001$.
3. To further examine which intervention method is most effective, we performed Tukey's post-hoc comparisons. The results revealed that all three intervention methods have significantly higher scores than the control group (G1-Ctrl: p = .003; G2-Ctrl: p \< .001; G3-Ctrl: p \< .001). Among the three intervention methods, G3 seems to be the most effective. Specifically, G3 showed significantly higher scores than G1 (p = .004). However, no significant difference was found between G2 and G3 (p = .105).
[Discussion]{.bigger}
These findings suggest that higher intervention intensity improves verbal acquisition performance, which is consistent with prior literature \[xxxx/references\].
# After-Class Exercise: Effect of Sleep Duration on Cognitive Performance
## Background
- Research Question:
- Does the amount of sleep affect cognitive performance on a standardized test?
- Study Design
- Independent variable: Sleep duration (3 groups: Short (≤5 hrs), Moderate (6-7 hrs), Long (≥8 hrs)).
- Dependent variable: Cognitive performance scores (measured as test scores out of 100).
---------------------------------------------
::: panel-tabset
### Exercise
```{webr-r}
#| context: setup
# Set seed for reproducibility
set.seed(42)
# Generate synthetic data for sleep study
sleep_data <- data.frame(
sleep_group = rep(c("Short", "Moderate", "Long"), each = 30),
cognitive_score = c(
rnorm(30, mean = 65, sd = 10), # Short sleep group (≤5 hrs)
rnorm(30, mean = 75, sd = 12), # Moderate sleep group (6-7 hrs)
rnorm(30, mean = 80, sd = 8) # Long sleep group (≥8 hrs)
)
)
```
```{webr-r}
# load packages you need
library(dplyr)
library(ggplot2)
library(lmtest)
library(car)
# View first few rows
head(sleep_data)
# Summary statistics by sleep group
sleep_data |>
group_by(sleep_group) |>
summarise(
_______
)
# Step 3: Check ANOVA Assumptions
# Indepdendence check: Perform the Durbin-Watson test
dw_test_result <- ____(____(_________, data = sleep_data))
# View the test results
print(dw_test_result)
# Normality check: Shapiro-Wilk normality test for each group
sleep_data |>
group_by(sleep_group) |>
summarise(
shapiro_p = ______(_______)$p.value
)
# HOV check: Levene's Test for homogeneity of variance
______(_______, data = sleep_data)
# Step 5: Perform One-Way ANOVA
anova_sleep <- ________(__________, data = sleep_data)
summary(anova_sleep)
# Step 6: Type I error control -- Tukey HSD post-hoc test
tukey_sleep <- TukeyHSD(_______)
tukey_sleep
```
### Answer
```{webr-r}
#| eval: false
#| echo: false
library(dplyr)
library(ggplot2)
library(lmtest)
library(car)
# View first few rows
head(sleep_data)
# Summary statistics by sleep group
sleep_data |>
group_by(sleep_group) |>
summarise(
mean_score = mean(cognitive_score),
sd_score = sd(cognitive_score),
n = n()
)
# Boxplot visualization
ggplot(sleep_data, aes(x = sleep_group, y = cognitive_score, fill = sleep_group)) +
geom_boxplot() +
theme_minimal() +
labs(title = "Cognitive Performance Across Sleep Groups", y = "Test Score", x = "Sleep Duration")
# Step 3: Check ANOVA Assumptions
# Indepdendence check: Perform the Durbin-Watson test
dw_test_result <- dwtest(aov(cognitive_score ~ sleep_group, data = sleep_data))
# View the test results
print(dw_test_result)
# Normality check: Shapiro-Wilk normality test for each group
sleep_data |>
group_by(sleep_group) |>
summarise(
shapiro_p = shapiro.test(cognitive_score)$p.value
)
ggplot(sleep_data, aes(sample = cognitive_score)) +
geom_qq() + geom_qq_line() +
facet_wrap(~sleep_group) +
theme_minimal() +
labs(title = "Q-Q Plot for Normality Check")
# HOV check: Levene's Test for homogeneity of variance
leveneTest(cognitive_score ~ sleep_group, data = sleep_data)
# Step 5: Perform One-Way ANOVA
anova_sleep <- aov(cognitive_score ~ sleep_group, data = sleep_data)
summary(anova_sleep)
# Step 6: Type I error control -- Tukey HSD post-hoc test
tukey_sleep <- TukeyHSD(anova_sleep)
tukey_sleep
```
::: heimu
- A one-way ANOVA was conducted to examine the effect of sleep duration on cognitive performance.
- There was a statistically significant difference in cognitive test scores across sleep groups, $F(2,87)=15.88$,$p<.001$.
- Tukey's post-hoc test revealed that participants in the Long sleep group (M=81.52, SD=6.27) performed significantly better than those in the Short sleep group (M=65.68, SD=12.55), p\<.01.
- These results suggest that inadequate sleep is associated with lower cognitive performance.
:::